DCN
Gene Information
- Official Symbol: DCN
- Official Name: decorin
- Aliases and Previous Symbols: N/A
- Entrez ID: 1634
- UniProt: P07585
- Interactions: BioGRID
- PubMed articles: Open PubMed
- OMIM: Open OMIM
Function Summary
- Entrez Summary: This gene encodes a member of the small leucine-rich proteoglycan family of proteins. Alternative splicing results in multiple transcript variants, at least one of which encodes a preproprotein that is proteolytically processed to generate the mature protein. This protein plays a role in collagen fibril assembly. Binding of this protein to multiple cell surface receptors mediates its role in tumor suppression, including a stimulatory effect on autophagy and inflammation and an inhibitory effect on angiogenesis and tumorigenesis. This gene and the related gene biglycan are thought to be the result of a gene duplication. Mutations in this gene are associated with congenital stromal corneal dystrophy in human patients. [provided by RefSeq, Nov 2015].
- UniProt Summary: May affect the rate of fibrils formation.
Pfam Domains GO Terms
Pfam Domains
| LRR 4 |
| LRRNT |
GO Terms
| collagen type VI trimer |
| peptide cross-linking via chondroitin 4-sulfate glycosaminoglycan |
| negative regulation of vascular endothelial growth factor signaling pathway |
| negative regulation of cellular response to vascular endothelial growth factor stimulus |
| positive regulation of mitochondrial depolarization |
| dermatan sulfate biosynthetic process |
| positive regulation of membrane depolarization |
| dermatan sulfate metabolic process |
| chondroitin sulfate catabolic process |
| positive regulation of mitochondrial fission |
| dermatan sulfate proteoglycan biosynthetic process |
| extracellular matrix structural constituent conferring compression resistance |
| dermatan sulfate proteoglycan metabolic process |
| regulation of vascular endothelial growth factor signaling pathway |
| regulation of cellular response to vascular endothelial growth factor stimulus |
| regulation of mitochondrial depolarization |
| regulation of mitochondrial fission |
| glycosaminoglycan binding |
| chondroitin sulfate biosynthetic process |
| extracellular matrix binding |
| chondroitin sulfate proteoglycan biosynthetic process |
| chondroitin sulfate metabolic process |
| regulation of membrane depolarization |
| chondroitin sulfate proteoglycan metabolic process |
| negative regulation of endothelial cell migration |
| sulfur compound catabolic process |
| peptide cross-linking |
| glycosaminoglycan catabolic process |
| collagen binding |
| negative regulation of epithelial cell migration |
| aminoglycan catabolic process |
| proteoglycan biosynthetic process |
| positive regulation of macroautophagy |
| regulation of mitochondrial membrane potential |
| positive regulation of phosphatidylinositol 3-kinase signaling |
| proteoglycan metabolic process |
| lysosomal lumen |
| Golgi lumen |
| negative regulation of angiogenesis |
| glycosaminoglycan biosynthetic process |
| negative regulation of blood vessel morphogenesis |
| mucopolysaccharide metabolic process |
| protein N-terminus binding |
| aminoglycan biosynthetic process |
| positive regulation of mitochondrion organization |
| negative regulation of vasculature development |
| regulation of phosphatidylinositol 3-kinase signaling |
| positive regulation of autophagy |
| skeletal muscle tissue development |
| skeletal muscle organ development |
| negative regulation of cellular response to growth factor stimulus |
| placenta development |
| glycosaminoglycan metabolic process |
| regulation of endothelial cell migration |
| aminoglycan metabolic process |
| regulation of macroautophagy |
| regulation of mitochondrion organization |
| sulfur compound biosynthetic process |
| monocarboxylic acid biosynthetic process |
| carbohydrate derivative catabolic process |
| peptidyl-serine modification |
| response to mechanical stimulus |
| regulation of epithelial cell migration |
| negative regulation of cell migration |
| kidney development |
| regulation of cellular response to growth factor stimulus |
| negative regulation of cell motility |
| aging |
| striated muscle tissue development |
| renal system development |
| regulation of angiogenesis |
| muscle organ development |
| muscle tissue development |
| carboxylic acid biosynthetic process |
| organic acid biosynthetic process |
| negative regulation of cellular component movement |
| response to lipopolysaccharide |
| urogenital system development |
| regulation of vasculature development |
| negative regulation of locomotion |
| glycoprotein biosynthetic process |
| response to molecule of bacterial origin |
| regulation of autophagy |
| extracellular matrix organization |
| collagen-containing extracellular matrix |
| positive regulation of cellular catabolic process |
| sulfur compound metabolic process |
| extracellular structure organization |
| glycoprotein metabolic process |
| reproductive structure development |
| reproductive system development |
| regulation of membrane potential |
| positive regulation of catabolic process |
| muscle structure development |
| wound healing |
| drug metabolic process |
| monocarboxylic acid metabolic process |
| response to wounding |
| small molecule biosynthetic process |
| carbohydrate derivative biosynthetic process |
| positive regulation of organelle organization |
| developmental process involved in reproduction |
| response to bacterium |
| ion homeostasis |
| regulation of cellular catabolic process |
| regulation of cell migration |
| response to lipid |
| peptidyl-amino acid modification |
| carboxylic acid metabolic process |
| regulation of cell motility |
| negative regulation of developmental process |
| animal organ morphogenesis |
| regulation of locomotion |
| regulation of catabolic process |
| regulation of cellular component movement |
| oxoacid metabolic process |
| positive regulation of intracellular signal transduction |
| organic acid metabolic process |
| carbohydrate derivative metabolic process |
| macromolecule catabolic process |
| organonitrogen compound catabolic process |
| regulation of anatomical structure morphogenesis |
| chemical homeostasis |
| response to abiotic stimulus |
| negative regulation of multicellular organismal process |
| positive regulation of cellular component organization |
| positive regulation of transcription by RNA polymerase II |
| negative regulation of signal transduction |
| regulation of organelle organization |
| response to other organism |
| response to external biotic stimulus |
| response to biotic stimulus |
| negative regulation of cell communication |
| negative regulation of signaling |
| positive regulation of developmental process |
| RNA binding |
| organonitrogen compound biosynthetic process |
| reproductive process |
| reproduction |
| positive regulation of transcription, DNA-templated |
| response to oxygen-containing compound |
| extracellular space |
| negative regulation of response to stimulus |
| positive regulation of nucleic acid-templated transcription |
| positive regulation of RNA biosynthetic process |
| homeostatic process |
| positive regulation of signal transduction |
| cellular macromolecule biosynthetic process |
| positive regulation of RNA metabolic process |
| small molecule metabolic process |
| tissue development |
| macromolecule biosynthetic process |
| organic substance catabolic process |
| cellular catabolic process |
| positive regulation of cell communication |
| positive regulation of signaling |
| regulation of intracellular signal transduction |
| positive regulation of nucleobase-containing compound metabolic process |
| positive regulation of macromolecule biosynthetic process |
| extracellular region |
| positive regulation of cellular biosynthetic process |
| positive regulation of gene expression |
| positive regulation of biosynthetic process |
CRISPR Data
Compound Hit Most Correlated Genes in Chemogenomics Tissues where Essential in the Avana Dataset (DepMap 20Q1)
Compound Hit
| Screen | Score |
|---|---|
| UM0011500 10μM R05 exp246 | -1.92 |
| VHL-ligand-1 20μM R07 exp421 | -1.87 |
| Dexamethasone 0.006μM R07 exp351 | -1.85 |
| CR131-b 0.005μM R08 exp474 | 1.91 |
Most Correlated Genes in Chemogenomics
No correlation found to any other genes in chemogenomics.
Tissues where Essential in the Avana Dataset (DepMap 20Q1)
Global Fraction of Cell Lines Where Essential: 0/739
| Tissue | Fraction Of Cell Lines Where Essential |
|---|---|
| 1290807.0 | 0/1 |
| 909776.0 | 0/1 |
| bile duct | 0/28 |
| blood | 0/28 |
| bone | 0/26 |
| breast | 0/33 |
| central nervous system | 0/56 |
| cervix | 0/4 |
| colorectal | 0/17 |
| esophagus | 0/13 |
| fibroblast | 0/1 |
| gastric | 0/16 |
| kidney | 0/21 |
| liver | 0/20 |
| lung | 0/75 |
| lymphocyte | 0/16 |
| ovary | 0/26 |
| pancreas | 0/24 |
| peripheral nervous system | 0/16 |
| plasma cell | 0/15 |
| prostate | 0/1 |
| skin | 0/24 |
| soft tissue | 0/9 |
| thyroid | 0/2 |
| upper aerodigestive | 0/22 |
| urinary tract | 0/29 |
| uterus | 0/5 |
Essentiality in NALM6
- Essentiality Rank: 13818
- Expression level (log2 read counts): 1.96
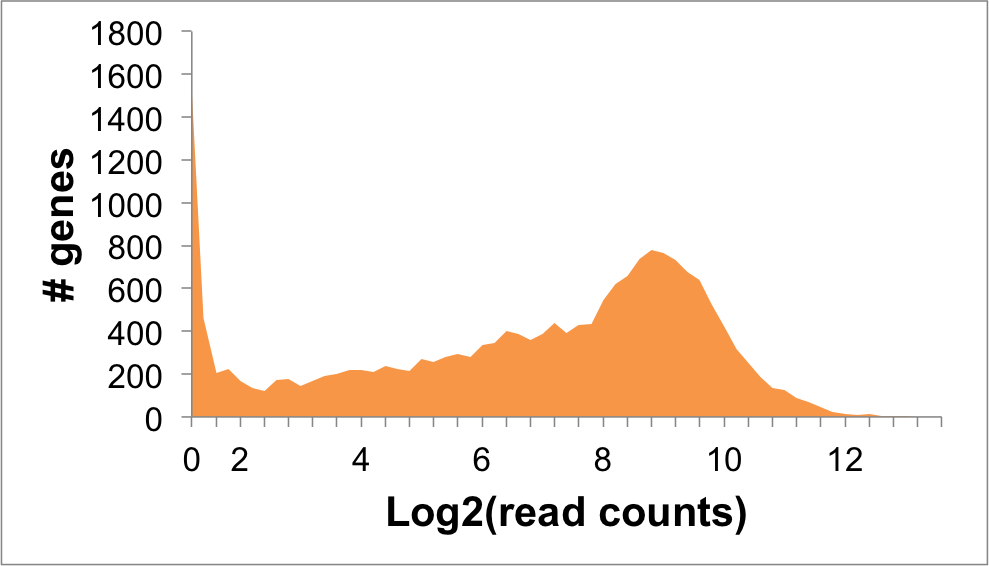
Expression Distribution
DCN Expression in NALM6 Cells: 1.96