NEDD4
Gene Information
- Official Symbol: NEDD4
- Official Name: NEDD4 E3 ubiquitin protein ligase
- Aliases and Previous Symbols: N/A
- Entrez ID: 4734
- UniProt: P46934
- Interactions: BioGRID
- PubMed articles: Open PubMed
- OMIM: Open OMIM
Function Summary
- Entrez Summary: This gene is the founding member of the NEDD4 family of HECT ubiquitin ligases that function in the ubiquitin proteasome system of protein degradation. The encoded protein contains an N-terminal calcium and phospholipid binding C2 domain followed by multiple tryptophan-rich WW domains and, a C-terminal HECT ubiquitin ligase catalytic domain. It plays critical role in the regulation of a number of membrane receptors, endocytic machinery components and the tumor suppressor PTEN. [provided by RefSeq, Jul 2016].
- UniProt Summary: N/A
Pfam Domains GO Terms
Pfam Domains
| WW |
| C2 |
| HECT |
GO Terms
| transmission of virus |
| dissemination or transmission of symbiont from host |
| development involved in symbiotic interaction |
| dissemination or transmission of organism from other organism involved in symbiotic interaction |
| negative regulation of transcription from RNA polymerase II promoter in response to UV-induced DNA damage |
| phosphothreonine residue binding |
| phosphoserine residue binding |
| regulation of transcription from RNA polymerase II promoter in response to UV-induced DNA damage |
| beta-2 adrenergic receptor binding |
| sodium channel inhibitor activity |
| glucocorticoid receptor signaling pathway |
| corticosteroid receptor signaling pathway |
| progesterone receptor signaling pathway |
| negative regulation of sodium ion transmembrane transporter activity |
| negative regulation of vascular endothelial growth factor receptor signaling pathway |
| negative regulation of sodium ion transmembrane transport |
| RNA polymerase binding |
| proline-rich region binding |
| negative regulation of sodium ion transport |
| apicolateral plasma membrane |
| protein targeting to lysosome |
| ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway |
| receptor catabolic process |
| regulation of vascular endothelial growth factor receptor signaling pathway |
| protein targeting to vacuole |
| protein localization to lysosome |
| protein K63-linked ubiquitination |
| neuromuscular junction development |
| establishment of protein localization to vacuole |
| regulation of sodium ion transmembrane transporter activity |
| regulation of potassium ion transmembrane transporter activity |
| receptor internalization |
| protein localization to vacuole |
| regulation of sodium ion transmembrane transport |
| positive regulation of nucleocytoplasmic transport |
| intracellular steroid hormone receptor signaling pathway |
| negative regulation of ion transmembrane transporter activity |
| negative regulation of transporter activity |
| ubiquitin binding |
| regulation of potassium ion transmembrane transport |
| negative regulation of cation transmembrane transport |
| cellular response to UV |
| positive regulation of phosphatidylinositol 3-kinase signaling |
| regulation of sodium ion transport |
| regulation of dendrite morphogenesis |
| negative regulation of ion transmembrane transport |
| regulation of potassium ion transport |
| receptor metabolic process |
| ubiquitin ligase complex |
| lysosomal transport |
| regulation of nucleocytoplasmic transport |
| cellular response to light stimulus |
| chromatin |
| negative regulation of transmembrane transport |
| regulation of phosphatidylinositol 3-kinase signaling |
| regulation of transcription from RNA polymerase II promoter in response to stress |
| steroid hormone mediated signaling pathway |
| regulation of DNA-templated transcription in response to stress |
| cell cortex |
| vacuolar transport |
| response to UV |
| negative regulation of cellular response to growth factor stimulus |
| negative regulation of ion transport |
| dendritic spine |
| regulation of dendrite development |
| response to calcium ion |
| interaction with host |
| intracellular receptor signaling pathway |
| hormone-mediated signaling pathway |
| regulation of macroautophagy |
| cellular response to radiation |
| cellular response to steroid hormone stimulus |
| positive regulation of intracellular transport |
| positive regulation of protein catabolic process |
| regulation of synapse organization |
| ubiquitin protein ligase activity |
| regulation of synapse structure or activity |
| receptor-mediated endocytosis |
| protein domain specific binding |
| regulation of ion transmembrane transporter activity |
| regulation of transmembrane transporter activity |
| synapse organization |
| regulation of cellular response to growth factor stimulus |
| regulation of transporter activity |
| protein polyubiquitination |
| regulation of cell morphogenesis involved in differentiation |
| response to light stimulus |
| cellular response to environmental stimulus |
| cellular response to abiotic stimulus |
| proteasome-mediated ubiquitin-dependent protein catabolic process |
| response to steroid hormone |
| regulation of autophagy |
| regulation of cation transmembrane transport |
| proteasomal protein catabolic process |
| regulation of intracellular transport |
| protein targeting |
| response to metal ion |
| regulation of protein catabolic process |
| regulation of metal ion transport |
| regulation of membrane potential |
| positive regulation of catabolic process |
| establishment of protein localization to organelle |
| response to radiation |
| regulation of ion transmembrane transport |
| negative regulation of transport |
| regulation of cell morphogenesis |
| regulation of neuron projection development |
| cellular response to lipid |
| ubiquitin-dependent protein catabolic process |
| modification-dependent protein catabolic process |
| response to inorganic substance |
| cellular response to organic cyclic compound |
| modification-dependent macromolecule catabolic process |
| endocytosis |
| regulation of transmembrane transport |
| proteolysis involved in cellular protein catabolic process |
| cellular response to hormone stimulus |
| cellular protein catabolic process |
| regulation of neuron differentiation |
| neuron projection development |
| import into cell |
| protein catabolic process |
| protein ubiquitination |
| regulation of plasma membrane bounded cell projection organization |
| perinuclear region of cytoplasm |
| regulation of ion transport |
| viral process |
| regulation of cell projection organization |
| protein localization to organelle |
| protein modification by small protein conjugation |
| cellular response to DNA damage stimulus |
| symbiotic process |
| neuron development |
| regulation of neurogenesis |
| interspecies interaction between organisms |
| regulation of cellular catabolic process |
| response to lipid |
| negative regulation of transcription by RNA polymerase II |
| cellular macromolecule catabolic process |
| response to hormone |
| regulation of cellular localization |
| response to organic cyclic compound |
| regulation of nervous system development |
| regulation of cell development |
| Golgi apparatus |
| protein modification by small protein conjugation or removal |
| positive regulation of transport |
| regulation of catabolic process |
| intracellular protein transport |
| neuron differentiation |
| positive regulation of intracellular signal transduction |
| macromolecule catabolic process |
| regulation of anatomical structure morphogenesis |
| organonitrogen compound catabolic process |
| plasma membrane bounded cell projection organization |
| negative regulation of molecular function |
| response to abiotic stimulus |
| cell projection organization |
| negative regulation of transcription, DNA-templated |
| cellular response to endogenous stimulus |
| negative regulation of nucleic acid-templated transcription |
| negative regulation of RNA biosynthetic process |
| negative regulation of signal transduction |
| proteolysis |
| locomotion |
| negative regulation of RNA metabolic process |
| negative regulation of cell communication |
| negative regulation of signaling |
| negative regulation of cellular macromolecule biosynthetic process |
| negative regulation of nucleobase-containing compound metabolic process |
| negative regulation of macromolecule biosynthetic process |
| response to endogenous stimulus |
| protein transport |
| negative regulation of cellular biosynthetic process |
| intracellular transport |
| generation of neurons |
| peptide transport |
| negative regulation of biosynthetic process |
| amide transport |
| cellular protein localization |
| cellular macromolecule localization |
| establishment of protein localization |
| negative regulation of response to stimulus |
| neurogenesis |
| cell development |
| positive regulation of signal transduction |
| cellular response to stress |
| positive regulation of protein metabolic process |
| negative regulation of gene expression |
| organic substance catabolic process |
| cellular catabolic process |
| regulation of cell differentiation |
| positive regulation of cell communication |
| positive regulation of signaling |
| regulation of intracellular signal transduction |
| establishment of localization in cell |
| nitrogen compound transport |
| regulation of transport |
| vesicle-mediated transport |
CRISPR Data
Compound Hit Most Correlated Genes in Chemogenomics Tissues where Essential in the Avana Dataset (DepMap 20Q1)
Compound Hit
| Screen | Score |
|---|---|
| IWR1 50μM R08 exp495 | 1.72 |
| Mdivi-1 15μM R05 exp217 | 1.96 |
Most Correlated Genes in Chemogenomics
| Gene | Correlation |
|---|---|
| PRIM1 | 0.407 |
Tissues where Essential in the Avana Dataset (DepMap 20Q1)
Global Fraction of Cell Lines Where Essential: 0/739
| Tissue | Fraction Of Cell Lines Where Essential |
|---|---|
| 1290807.0 | 0/1 |
| 909776.0 | 0/1 |
| bile duct | 0/28 |
| blood | 0/28 |
| bone | 0/26 |
| breast | 0/33 |
| central nervous system | 0/56 |
| cervix | 0/4 |
| colorectal | 0/17 |
| esophagus | 0/13 |
| fibroblast | 0/1 |
| gastric | 0/16 |
| kidney | 0/21 |
| liver | 0/20 |
| lung | 0/75 |
| lymphocyte | 0/16 |
| ovary | 0/26 |
| pancreas | 0/24 |
| peripheral nervous system | 0/16 |
| plasma cell | 0/15 |
| prostate | 0/1 |
| skin | 0/24 |
| soft tissue | 0/9 |
| thyroid | 0/2 |
| upper aerodigestive | 0/22 |
| urinary tract | 0/29 |
| uterus | 0/5 |
Essentiality in NALM6
- Essentiality Rank: 2886
- Expression level (log2 read counts): 6.29
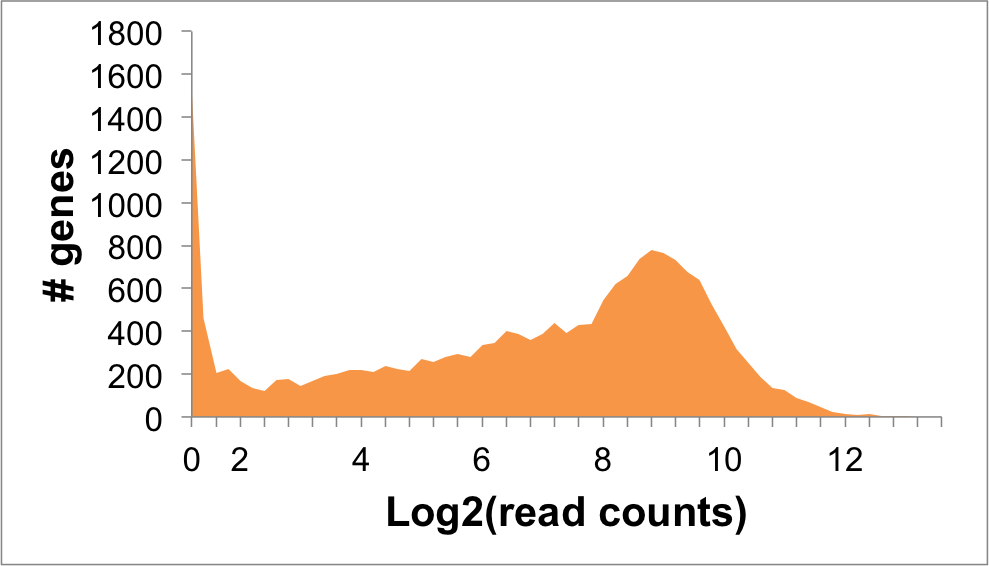
Expression Distribution
NEDD4 Expression in NALM6 Cells: 6.29