CCL2
Gene Information
- Official Symbol: CCL2
- Official Name: C-C motif chemokine ligand 2
- Aliases and Previous Symbols: N/A
- Entrez ID: 6347
- UniProt: P13500
- Interactions: BioGRID
- PubMed articles: Open PubMed
- OMIM: Open OMIM
Function Summary
- Entrez Summary: This gene is one of several cytokine genes clustered on the q-arm of chromosome 17. Chemokines are a superfamily of secreted proteins involved in immunoregulatory and inflammatory processes. The superfamily is divided into four subfamilies based on the arrangement of N-terminal cysteine residues of the mature peptide. This chemokine is a member of the CC subfamily which is characterized by two adjacent cysteine residues. This cytokine displays chemotactic activity for monocytes and basophils but not for neutrophils or eosinophils. It has been implicated in the pathogenesis of diseases characterized by monocytic infiltrates, like psoriasis, rheumatoid arthritis and atherosclerosis. It binds to chemokine receptors CCR2 and CCR4. [provided by RefSeq, Jul 2013].
- UniProt Summary: Chemotactic factor that attracts monocytes and basophils but not neutrophils or eosinophils. Augments monocyte anti-tumor activity. Has been implicated in the pathogenesis of diseases characterized by monocytic infiltrates, like psoriasis, rheumatoid arthritis or atherosclerosis. May be involved in the recruitment of monocytes into the arterial wall during the disease process of atherosclerosis.
Pfam Domains GO Terms
Pfam Domains
| IL8 |
GO Terms
| negative regulation of natural killer cell chemotaxis |
| helper T cell extravasation |
| negative regulation of lymphocyte chemotaxis |
| CCR2 chemokine receptor binding |
| negative regulation of vascular endothelial cell proliferation |
| astrocyte cell migration |
| positive regulation of NMDA glutamate receptor activity |
| T cell extravasation |
| regulation of natural killer cell chemotaxis |
| negative regulation of glial cell apoptotic process |
| positive regulation of apoptotic cell clearance |
| regulation of apoptotic cell clearance |
| regulation of glial cell apoptotic process |
| PERK-mediated unfolded protein response |
| macrophage chemotaxis |
| negative regulation of lymphocyte migration |
| positive regulation of nitric-oxide synthase biosynthetic process |
| positive regulation of glutamate receptor signaling pathway |
| negative regulation of leukocyte chemotaxis |
| macrophage migration |
| positive regulation of endothelial cell apoptotic process |
| regulation of nitric-oxide synthase biosynthetic process |
| regulation of vascular endothelial cell proliferation |
| positive regulation of calcium ion import |
| eosinophil chemotaxis |
| eosinophil migration |
| regulation of lymphocyte chemotaxis |
| T cell migration |
| viral genome replication |
| CCR chemokine receptor binding |
| positive regulation of synaptic transmission, glutamatergic |
| positive regulation of epithelial cell apoptotic process |
| positive regulation of signaling receptor activity |
| lipopolysaccharide-mediated signaling pathway |
| ER-nucleus signaling pathway |
| regulation of NMDA receptor activity |
| glial cell migration |
| negative regulation of endothelial cell proliferation |
| receptor signaling pathway via JAK-STAT |
| protein kinase B signaling |
| regulation of calcium ion import |
| receptor signaling pathway via STAT |
| cellular extravasation |
| monocyte chemotaxis |
| negative regulation of leukocyte migration |
| lymphocyte chemotaxis |
| mononuclear cell migration |
| chemokine activity |
| regulation of endothelial cell apoptotic process |
| negative regulation of chemotaxis |
| regulation of lymphocyte migration |
| regulation of glutamate receptor signaling pathway |
| lymphocyte migration |
| positive regulation of phagocytosis |
| regulation of synaptic transmission, glutamatergic |
| positive regulation of cation channel activity |
| sensory perception of pain |
| regulation of neurotransmitter receptor activity |
| regulation of epithelial cell apoptotic process |
| chemokine-mediated signaling pathway |
| cell |
| neutrophil chemotaxis |
| granulocyte chemotaxis |
| cellular response to chemokine |
| response to chemokine |
| neutrophil migration |
| regulation of phagocytosis |
| granulocyte migration |
| negative regulation of G1/S transition of mitotic cell cycle |
| positive regulation of ion transmembrane transporter activity |
| endoplasmic reticulum unfolded protein response |
| negative regulation of cell cycle G1/S phase transition |
| cellular response to fibroblast growth factor stimulus |
| positive regulation of transporter activity |
| regulation of leukocyte chemotaxis |
| response to fibroblast growth factor |
| positive regulation of calcium ion transport |
| negative regulation of epithelial cell proliferation |
| cellular response to unfolded protein |
| myeloid leukocyte migration |
| regulation of endothelial cell proliferation |
| positive regulation of cation transmembrane transport |
| negative regulation of neuron apoptotic process |
| leukocyte chemotaxis |
| cellular response to topologically incorrect protein |
| regulation of G1/S transition of mitotic cell cycle |
| positive regulation of synaptic transmission |
| regulation of cell shape |
| positive regulation of ion transmembrane transport |
| cellular response to interferon-gamma |
| response to unfolded protein |
| regulation of cell cycle G1/S phase transition |
| regulation of cation channel activity |
| regulation of signaling receptor activity |
| cellular response to interleukin-1 |
| response to interferon-gamma |
| cellular response to lipopolysaccharide |
| response to topologically incorrect protein |
| cellular response to molecule of bacterial origin |
| regulation of leukocyte migration |
| response to interleukin-1 |
| viral life cycle |
| positive regulation of T cell activation |
| positive regulation of transmembrane transport |
| negative regulation of neuron death |
| regulation of neuron apoptotic process |
| cell chemotaxis |
| positive regulation of ERK1 and ERK2 cascade |
| regulation of chemotaxis |
| negative regulation of mitotic cell cycle phase transition |
| gliogenesis |
| cellular response to biotic stimulus |
| positive regulation of leukocyte cell-cell adhesion |
| negative regulation of cell cycle phase transition |
| protein kinase activity |
| cellular response to tumor necrosis factor |
| G protein-coupled receptor signaling pathway, coupled to cyclic nucleotide second messenger |
| regulation of calcium ion transport |
| positive regulation of cell-cell adhesion |
| response to endoplasmic reticulum stress |
| regulation of ion transmembrane transporter activity |
| negative regulation of cell migration |
| response to tumor necrosis factor |
| regulation of transmembrane transporter activity |
| negative regulation of cell motility |
| positive regulation of ion transport |
| regulation of transporter activity |
| regulation of ERK1 and ERK2 cascade |
| regulation of leukocyte cell-cell adhesion |
| negative regulation of mitotic cell cycle |
| negative regulation of cellular component movement |
| regulation of neuron death |
| response to lipopolysaccharide |
| angiogenesis |
| regulation of T cell activation |
| negative regulation of locomotion |
| negative regulation of cell cycle process |
| response to molecule of bacterial origin |
| regulation of cation transmembrane transport |
| regulation of epithelial cell proliferation |
| signaling receptor binding |
| humoral immune response |
| positive regulation of lymphocyte activation |
| negative regulation of response to external stimulus |
| MAPK cascade |
| leukocyte migration |
| regulation of metal ion transport |
| signal transduction by protein phosphorylation |
| regulation of cell-cell adhesion |
| positive regulation of cell adhesion |
| positive regulation of leukocyte activation |
| blood vessel morphogenesis |
| regulation of mitotic cell cycle phase transition |
| positive regulation of GTPase activity |
| positive regulation of cell activation |
| negative regulation of immune system process |
| modulation of chemical synaptic transmission |
| regulation of trans-synaptic signaling |
| regulation of cell cycle phase transition |
| regulation of ion transmembrane transport |
| regulation of GTPase activity |
| blood vessel development |
| regulation of cell morphogenesis |
| inflammatory response |
| cellular response to growth factor stimulus |
| vasculature development |
| regulation of lymphocyte activation |
| cardiovascular system development |
| cellular response to lipid |
| response to growth factor |
| cellular response to organic cyclic compound |
| positive regulation of MAPK cascade |
| regulation of vesicle-mediated transport |
| chemotaxis |
| taxis |
| regulation of transmembrane transport |
| negative regulation of cell cycle |
| regulation of leukocyte activation |
| regulation of mitotic cell cycle |
| regulation of cell activation |
| positive regulation of apoptotic process |
| positive regulation of programmed cell death |
| tube morphogenesis |
| cytokine-mediated signaling pathway |
| negative regulation of cell population proliferation |
| regulation of cell adhesion |
| response to bacterium |
| positive regulation of cell death |
| regulation of ion transport |
| viral process |
| regulation of MAPK cascade |
| regulation of cell cycle process |
| innate immune response |
| positive regulation of hydrolase activity |
| symbiotic process |
| interspecies interaction between organisms |
| tube development |
| regulation of cell migration |
| response to lipid |
| circulatory system development |
| negative regulation of apoptotic process |
| anatomical structure formation involved in morphogenesis |
| cellular homeostasis |
| negative regulation of programmed cell death |
| regulation of cell motility |
| response to organic cyclic compound |
| cell adhesion |
| biological adhesion |
| defense response to other organism |
| animal organ morphogenesis |
| cell migration |
| sensory perception |
| protein phosphorylation |
| regulation of locomotion |
| positive regulation of transport |
| negative regulation of cell death |
| regulation of cellular component movement |
| cellular response to cytokine stimulus |
| positive regulation of protein phosphorylation |
| positive regulation of intracellular signal transduction |
| cellular response to oxygen-containing compound |
| positive regulation of phosphorylation |
| regulation of anatomical structure morphogenesis |
| cell motility |
| localization of cell |
| regulation of response to external stimulus |
| response to cytokine |
| cytoskeleton organization |
| positive regulation of phosphorus metabolic process |
| positive regulation of phosphate metabolic process |
| positive regulation of immune system process |
| regulation of cell cycle |
| cellular response to endogenous stimulus |
| positive regulation of protein modification process |
| regulation of hydrolase activity |
| phosphorylation |
| response to other organism |
| response to external biotic stimulus |
| locomotion |
| G protein-coupled receptor signaling pathway |
| response to biotic stimulus |
| defense response |
| nervous system process |
| positive regulation of catalytic activity |
| regulation of protein phosphorylation |
| response to endogenous stimulus |
| regulation of apoptotic process |
| movement of cell or subcellular component |
| response to oxygen-containing compound |
| regulation of programmed cell death |
| regulation of phosphorylation |
| extracellular space |
| positive regulation of cellular protein metabolic process |
| regulation of cell population proliferation |
| negative regulation of response to stimulus |
| neurogenesis |
| homeostatic process |
| regulation of immune system process |
| positive regulation of signal transduction |
| regulation of cell death |
| intracellular signal transduction |
| cellular response to stress |
| positive regulation of protein metabolic process |
| positive regulation of molecular function |
| regulation of phosphate metabolic process |
| regulation of phosphorus metabolic process |
| positive regulation of cell communication |
| positive regulation of signaling |
| regulation of intracellular signal transduction |
| regulation of protein modification process |
| regulation of transport |
| immune response |
| positive regulation of macromolecule biosynthetic process |
| extracellular region |
| system process |
| positive regulation of biosynthetic process |
CRISPR Data
Compound Hit Most Correlated Genes in Chemogenomics Tissues where Essential in the Avana Dataset (DepMap 20Q1)
Compound Hit
| Screen | Score |
|---|---|
| Trientine 500μM R08 exp534 | -1.95 |
| UM0125461 0.74μM R02 exp84 | -1.79 |
| UM0011500 10μM R05 exp246 | -1.71 |
| Diepoxybutane 3μM R07 exp354 | 2.03 |
| Cidofovir 10μM R07 exp345 | 2.22 |
Most Correlated Genes in Chemogenomics
No correlation found to any other genes in chemogenomics.
Tissues where Essential in the Avana Dataset (DepMap 20Q1)
Global Fraction of Cell Lines Where Essential: 0/739
| Tissue | Fraction Of Cell Lines Where Essential |
|---|---|
| 1290807.0 | 0/1 |
| 909776.0 | 0/1 |
| bile duct | 0/28 |
| blood | 0/28 |
| bone | 0/26 |
| breast | 0/33 |
| central nervous system | 0/56 |
| cervix | 0/4 |
| colorectal | 0/17 |
| esophagus | 0/13 |
| fibroblast | 0/1 |
| gastric | 0/16 |
| kidney | 0/21 |
| liver | 0/20 |
| lung | 0/75 |
| lymphocyte | 0/16 |
| ovary | 0/26 |
| pancreas | 0/24 |
| peripheral nervous system | 0/16 |
| plasma cell | 0/15 |
| prostate | 0/1 |
| skin | 0/24 |
| soft tissue | 0/9 |
| thyroid | 0/2 |
| upper aerodigestive | 0/22 |
| urinary tract | 0/29 |
| uterus | 0/5 |
Essentiality in NALM6
- Essentiality Rank: 14983
- Expression level (log2 read counts): -7.68
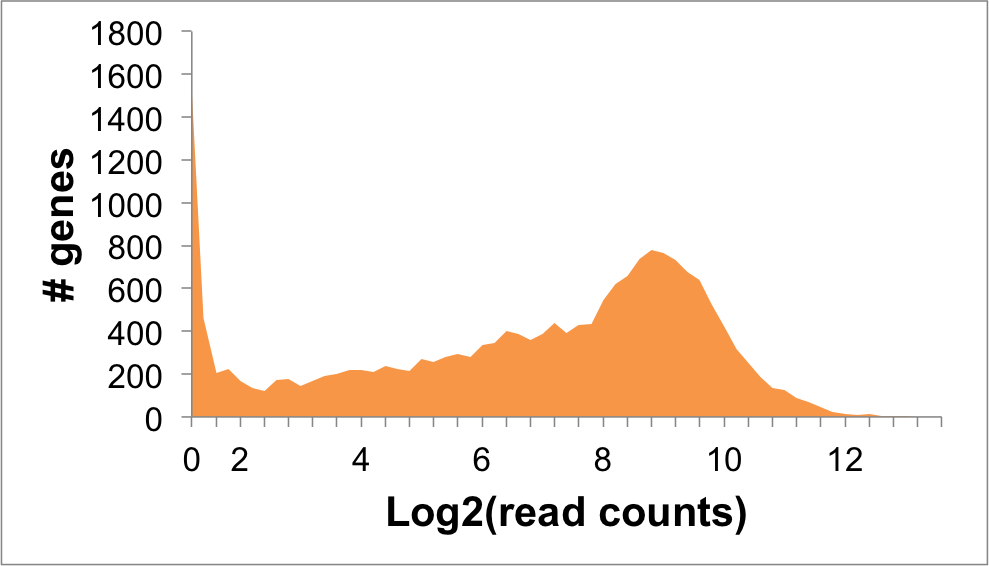
Expression Distribution
CCL2 Expression in NALM6 Cells: -7.68