TDP2
Gene Information
- Official Symbol: TDP2
- Official Name: tyrosyl-DNA phosphodiesterase 2
- Aliases and Previous Symbols: N/A
- Entrez ID: 51567
- UniProt: O95551
- Interactions: BioGRID
- PubMed articles: Open PubMed
- OMIM: Open OMIM
Function Summary
- Entrez Summary: This gene encodes a member of a superfamily of divalent cation-dependent phosphodiesterases. The encoded protein associates with CD40, tumor necrosis factor (TNF) receptor-75 and TNF receptor associated factors (TRAFs), and inhibits nuclear factor-kappa-B activation. This protein has sequence and structural similarities with APE1 endonuclease, which is involved in both DNA repair and the activation of transcription factors. [provided by RefSeq, Jul 2008].
- UniProt Summary: DNA repair enzyme that can remove a variety of covalent adducts from DNA through hydrolysis of a 5'-phosphodiester bond, giving rise to DNA with a free 5' phosphate. Catalyzes the hydrolysis of dead-end complexes between DNA and the topoisomerase 2 (TOP2) active site tyrosine residue. The 5'-tyrosyl DNA phosphodiesterase activity can enable the repair of TOP2-induced DNA double-strand breaks/DSBs without the need for nuclease activity, creating a 'clean' DSB with 5'-phosphate termini that are ready for ligation. Thereby, protects the transcription of many genes involved in neurological development and maintenance from the abortive activity of TOP2. Hydrolyzes 5'- phosphoglycolates on protruding 5' ends on DSBs due to DNA damage by radiation and free radicals. Has preference for single-stranded DNA or duplex DNA with a 4 base pair overhang as substrate. Acts as a regulator of ribosome biogenesis following stress. Has also 3'-tyrosyl DNA phosphodiesterase activity, but less efficiently and much slower than TDP1. Constitutes the major if not only 5'- tyrosyl-DNA phosphodiesterase in cells. Also acts as an adapter by participating in the specific activation of MAP3K7/TAK1 in response to TGF-beta: associates with components of the TGF-beta receptor-TRAF6-TAK1 signaling module and promotes their ubiquitination dependent complex formation. Involved in non- canonical TGF-beta induced signaling routes. May also act as a negative regulator of ETS1 and may inhibit NF-kappa-B activation. {ECO:0000269|PubMed:19794497, ECO:0000269|PubMed:21030584, ECO:0000269|PubMed:21921940, ECO:0000269|PubMed:21980489, ECO:0000269|PubMed:22405347, ECO:0000269|PubMed:22822062, ECO:0000269|PubMed:24658003}.
CRISPR Data
Essentiality in NALM6
- Essentiality Rank: 3494
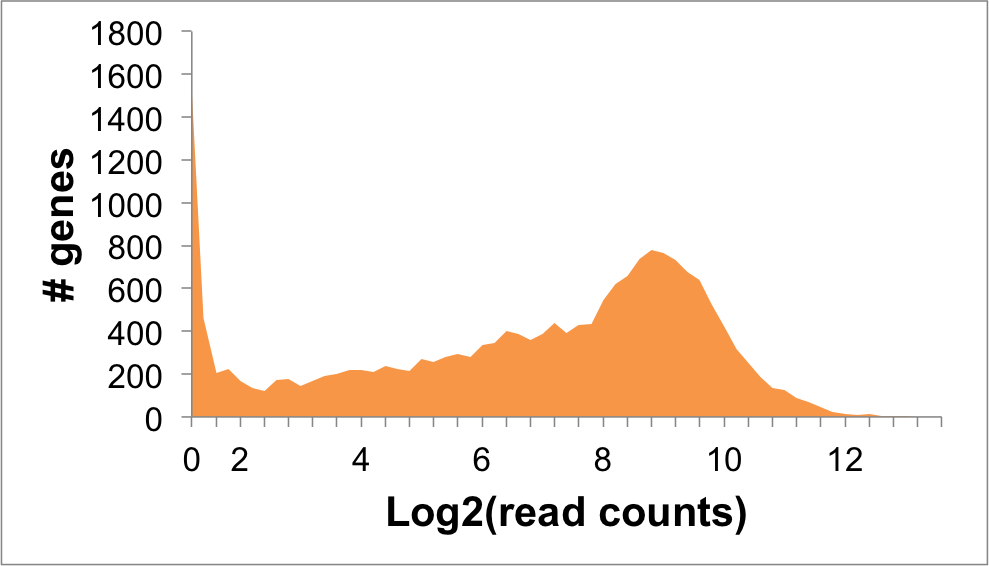
- Expression level (log2 read counts): 5.65