Table of Contents
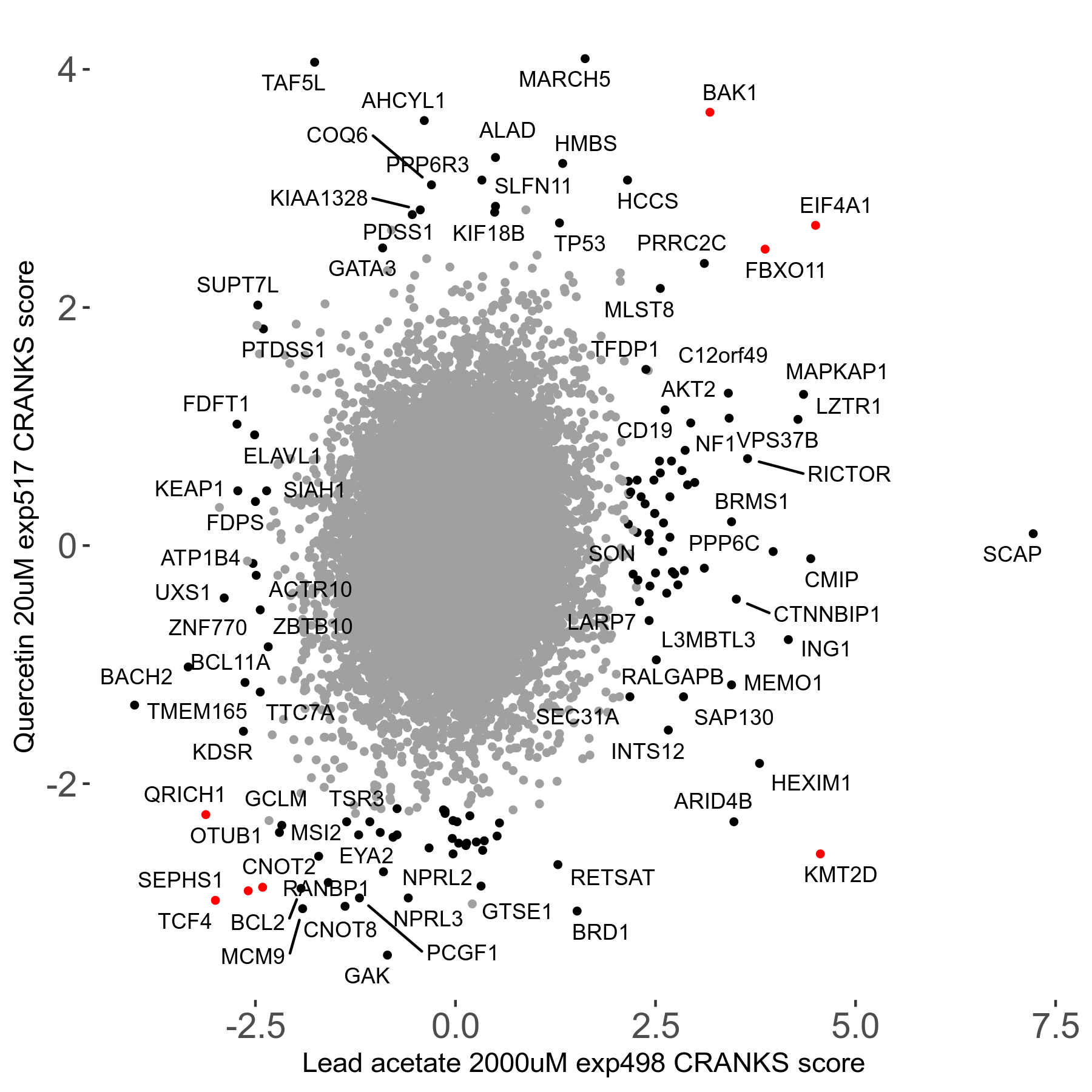
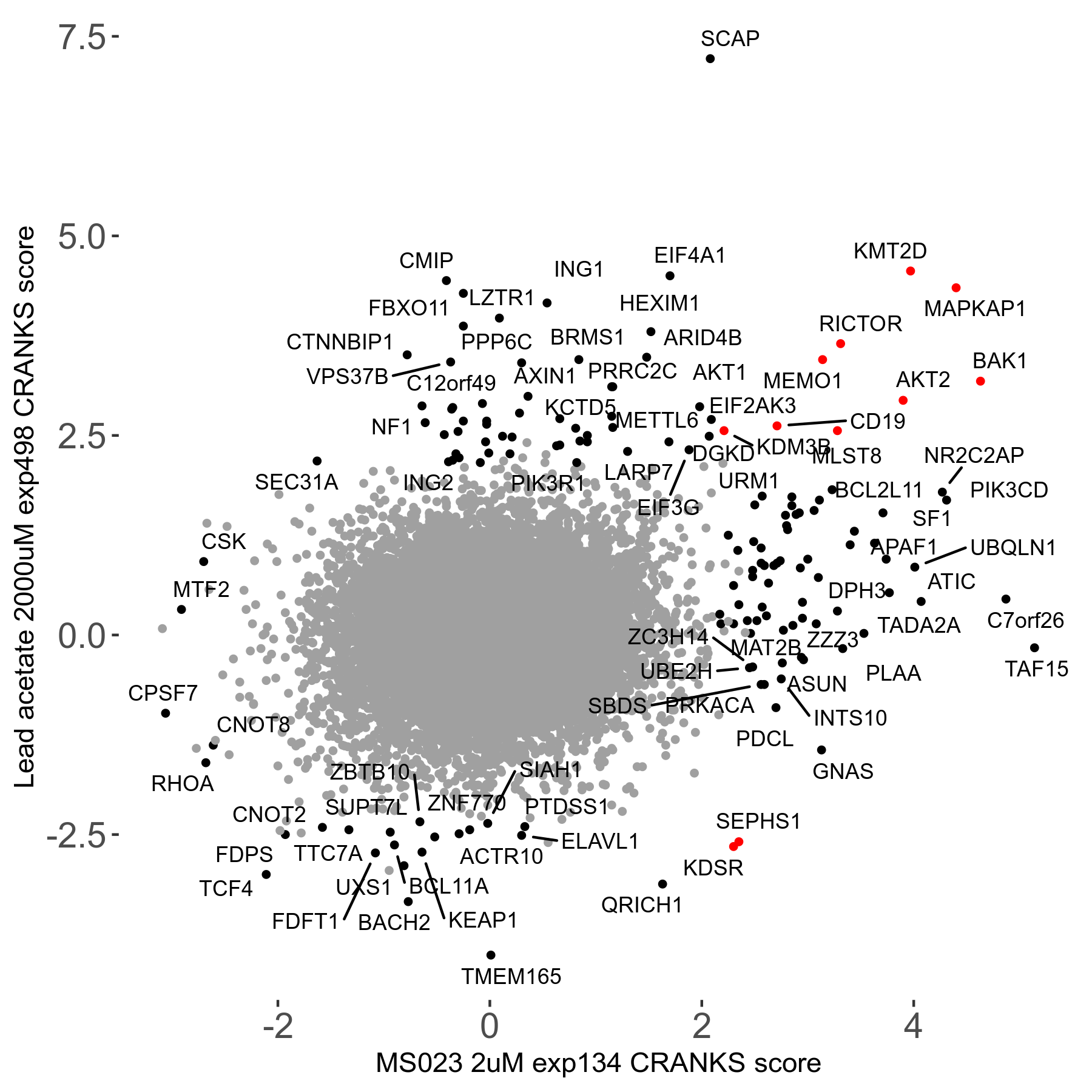
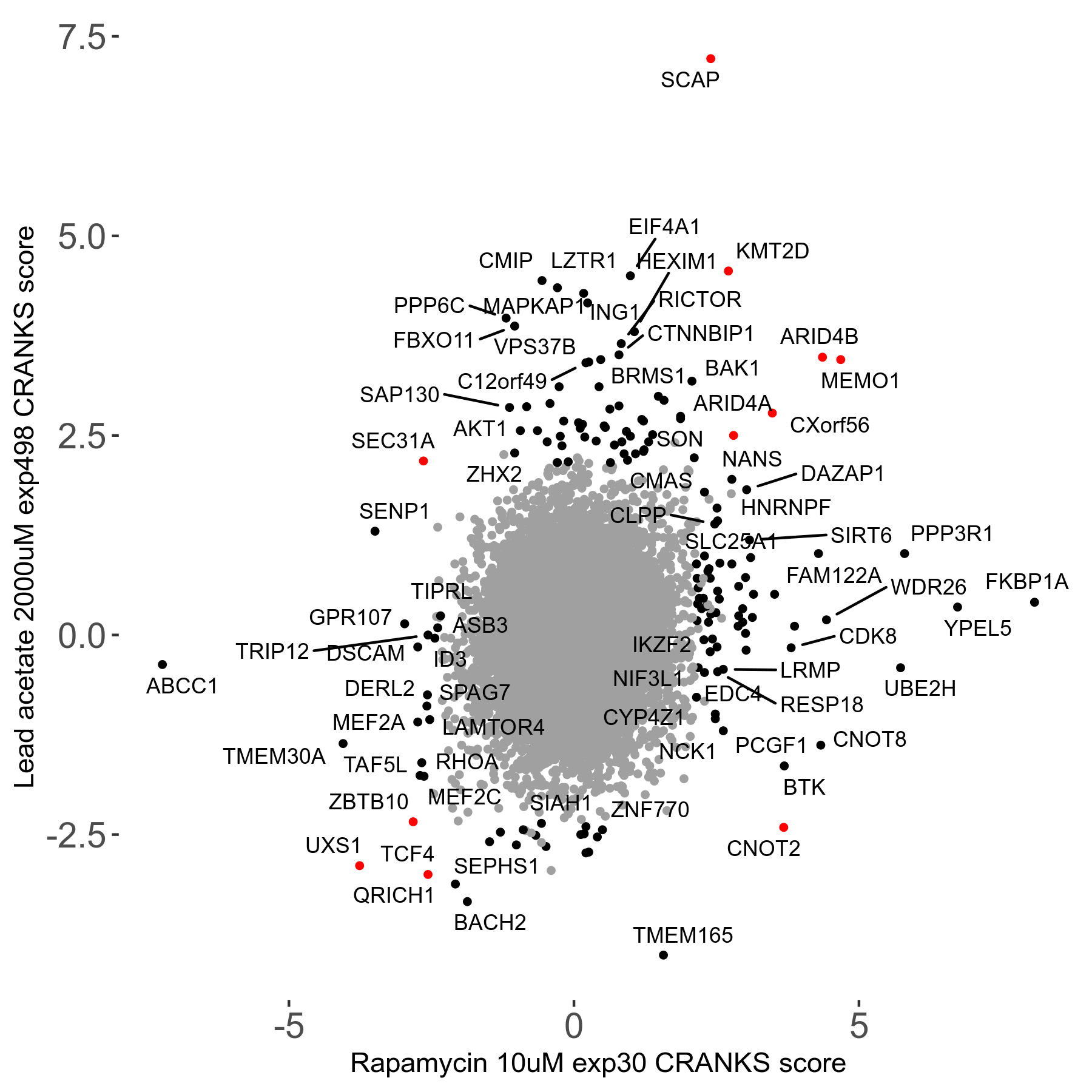
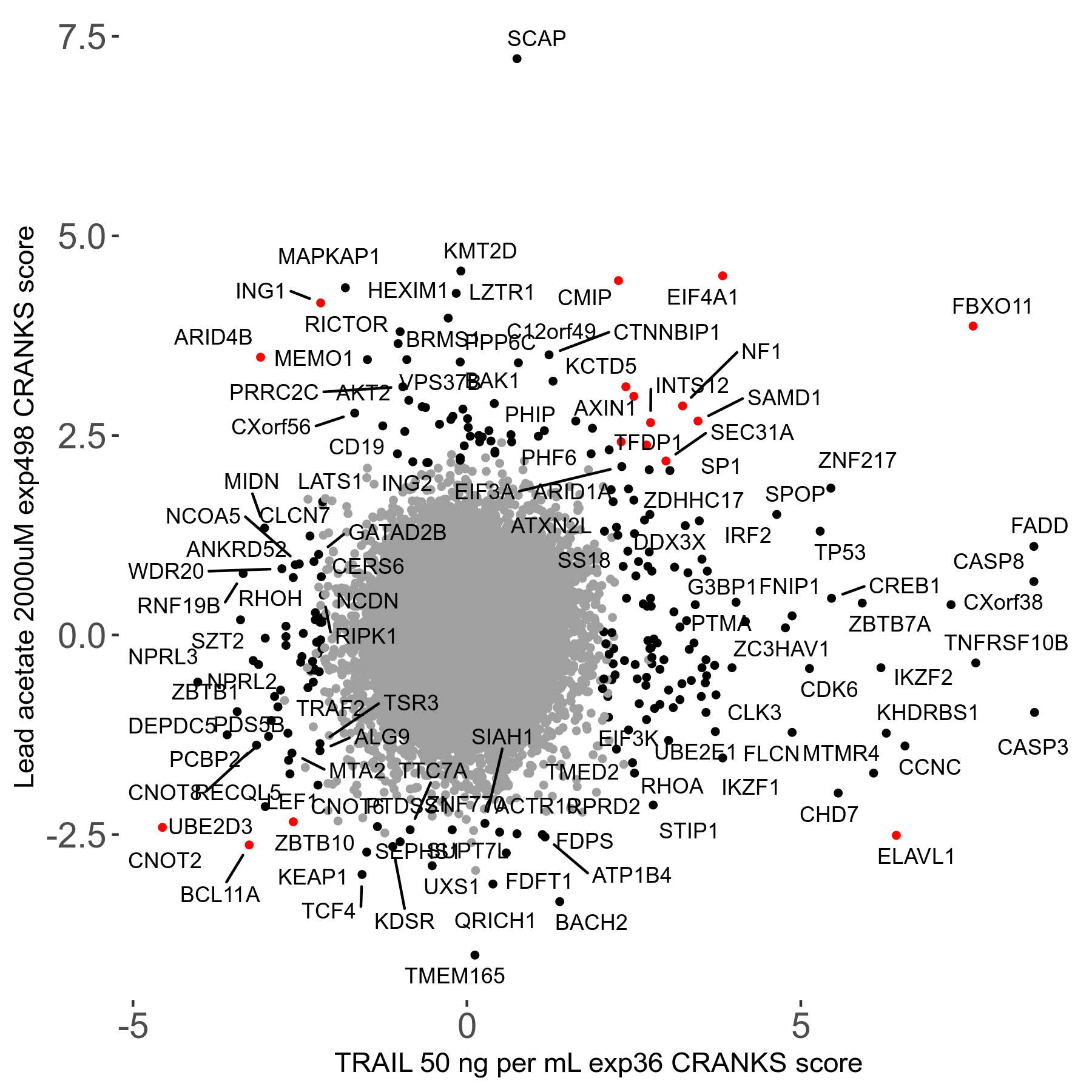
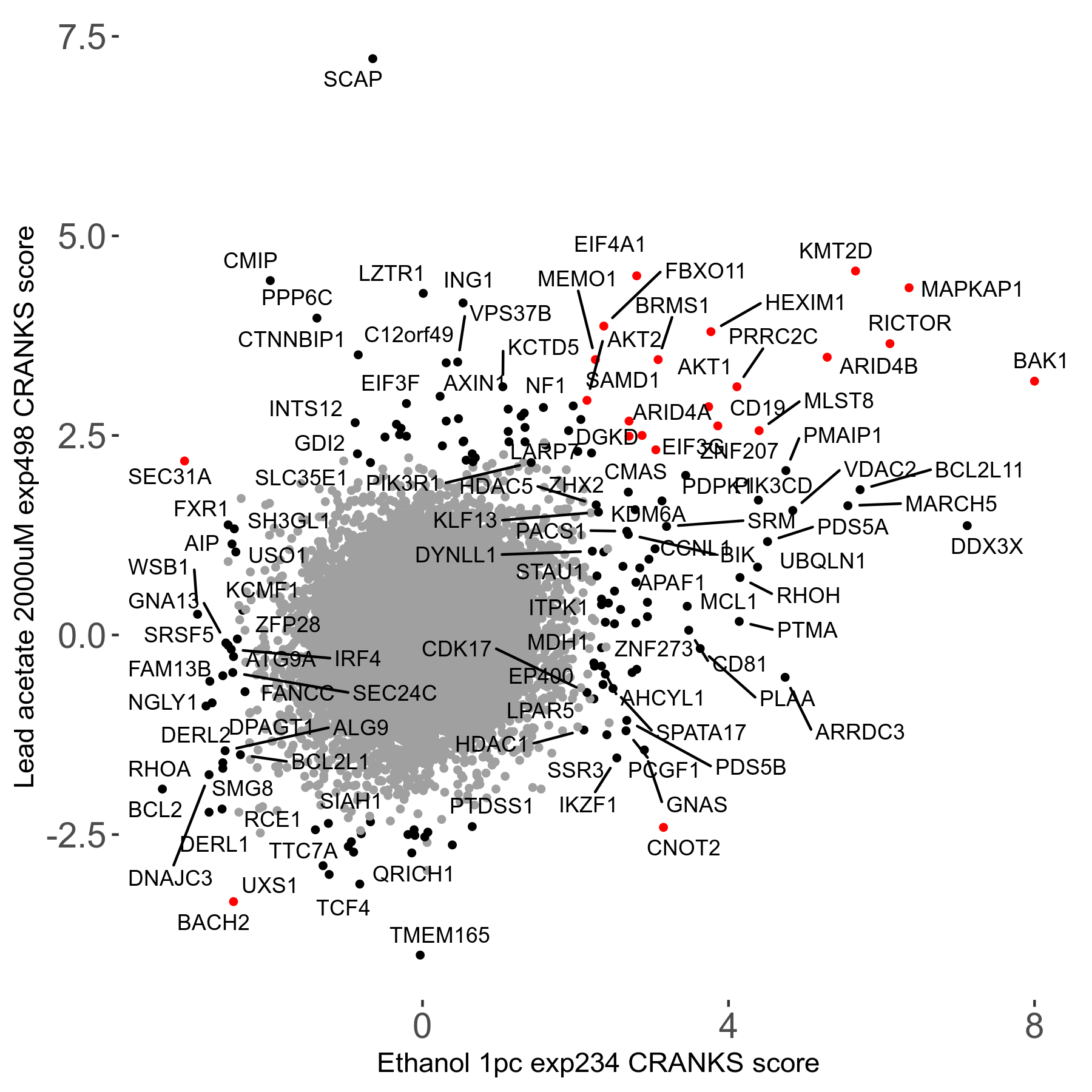
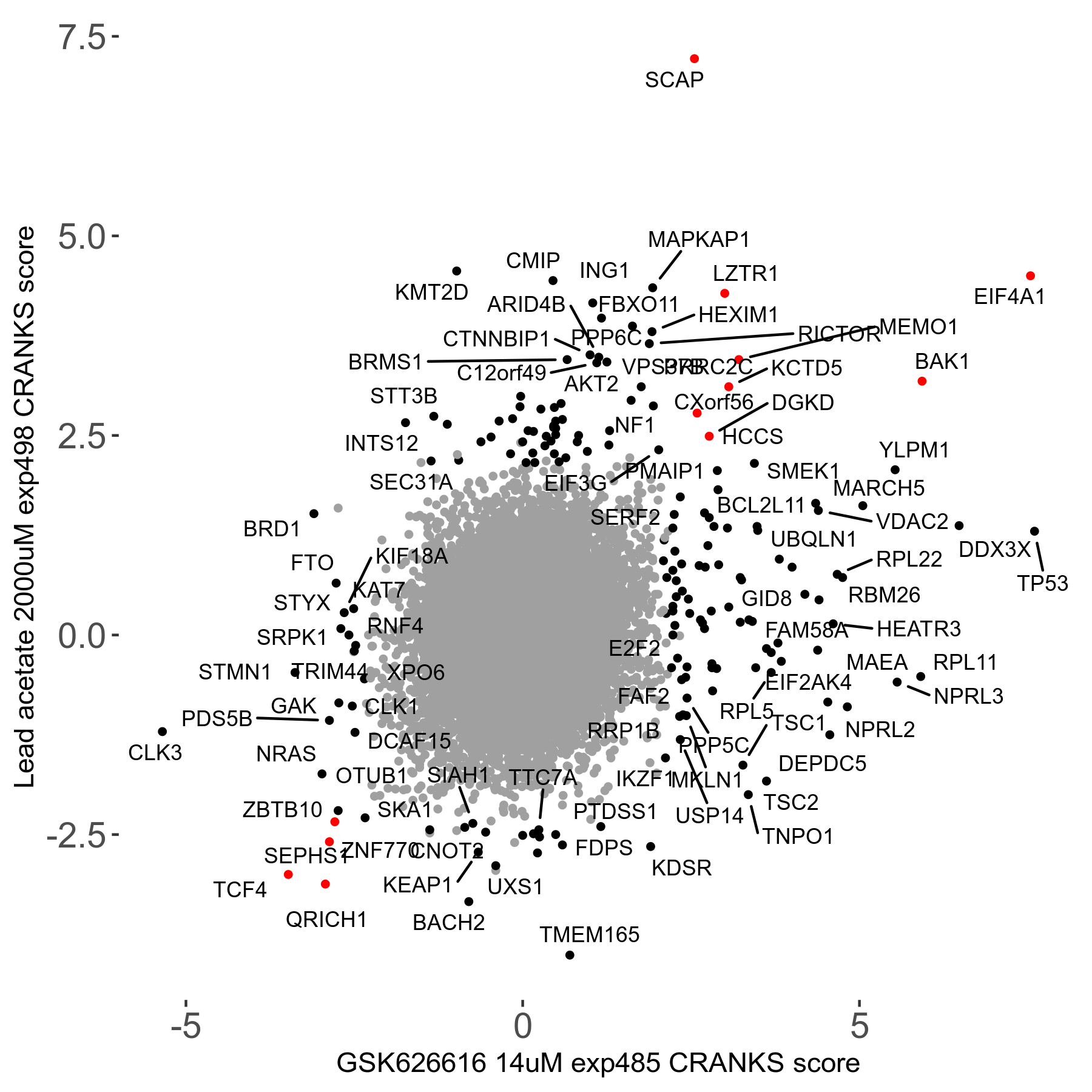
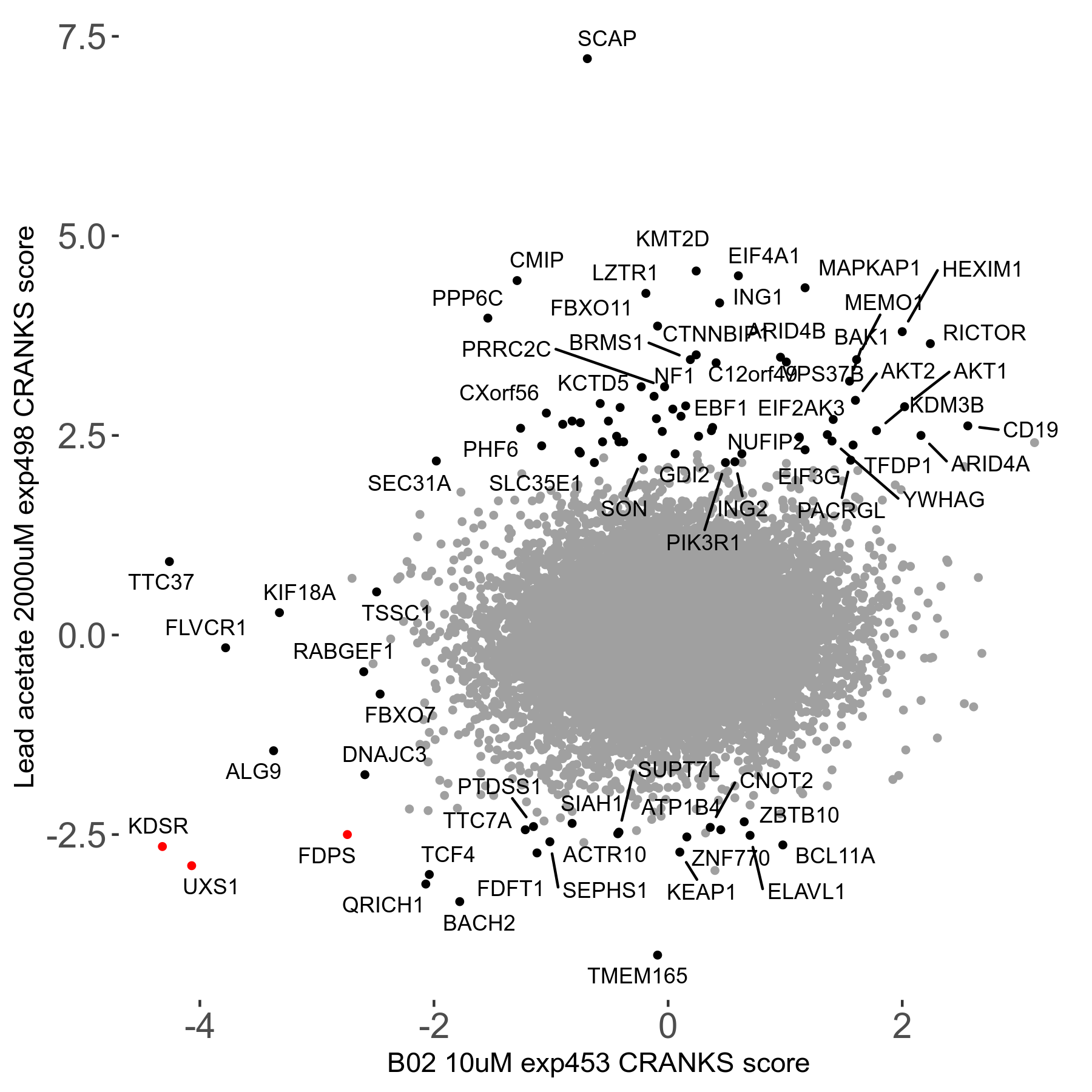
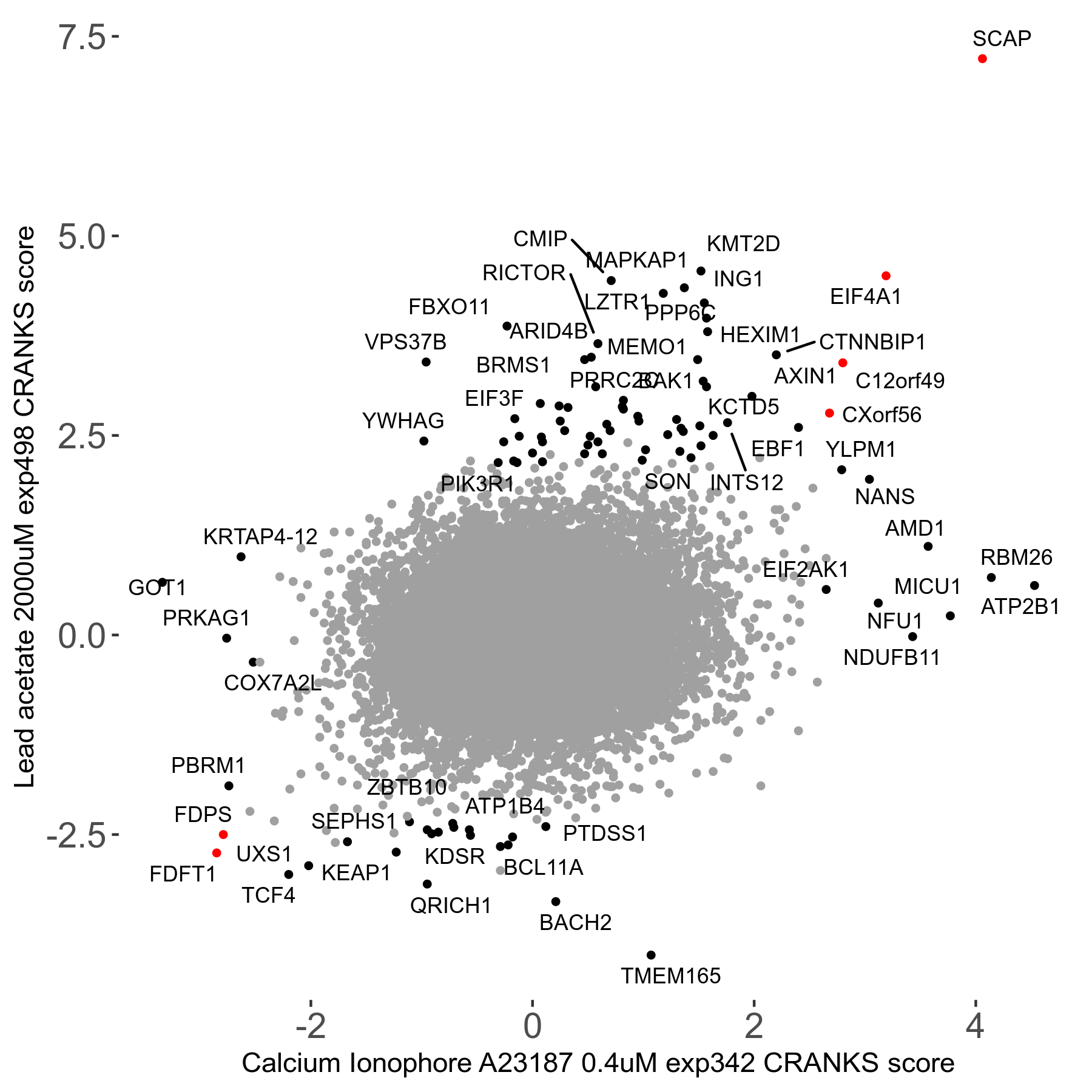
Lead acetate 2000μM R08 exp498 no dilution day6
Mechanism of Action
Industrial pollutant, heavy metal toxin
- Class / Subclass 1: Environmental Stresses / Heavy Metal
Technical Notes
Compound References
- PubChem Name: Lead acetate trihydrate
- Synonyms: N/A
- CAS #: 6080-56-4
- PubChem CID: 22456
- IUPAC: lead(2+);diacetate;trihydrate
- INCHI Name: InChI=1S/2C2H4O2.3H2O.Pb/c2*1-2(3)4;;;;/h2*1H3,(H,3,4);3*1H2;/q;;;;;+2/p-2
- INCHI Key: MCEUZMYFCCOOQO-UHFFFAOYSA-L
- Molecular Weight: 379
- Canonical SMILES: CC(=O)[O-].CC(=O)[O-].O.O.O.[Pb+2]
- Isomeric SMILES: N/A
- Molecular Formula: C4H12O7Pb
Compound Supplier
- Supplier Name: EM Science
- Catalog #: LX0120-1
- Lot #: 28147208
Compound Characterization
- XRPD: Expected Pb(CH3COO)2 3H2O; found Pb(CH3COO)2 +
Pb(CH3COO)2 3H2O
Dose Response Curve
- Platform ID: Lead-Aceteate
- Min: 2.9017; Max: 98.7157

| IC | Concentration (µM) |
|---|---|
| IC10 | N/A |
| IC20 | 1.3598 |
| IC30 | 2.1266 |
| IC40 | 3.0284 |
| IC50 | 4.1663 |
| IC60 | N/A |
| IC70 | N/A |
| IC80 | N/A |
| IC90 | N/A |
Screen Summary
- Round: 08
- Dose: 2000µM
- Days of incubation: 8
- Doublings: 2.0
- Numbers of reads: 16475946
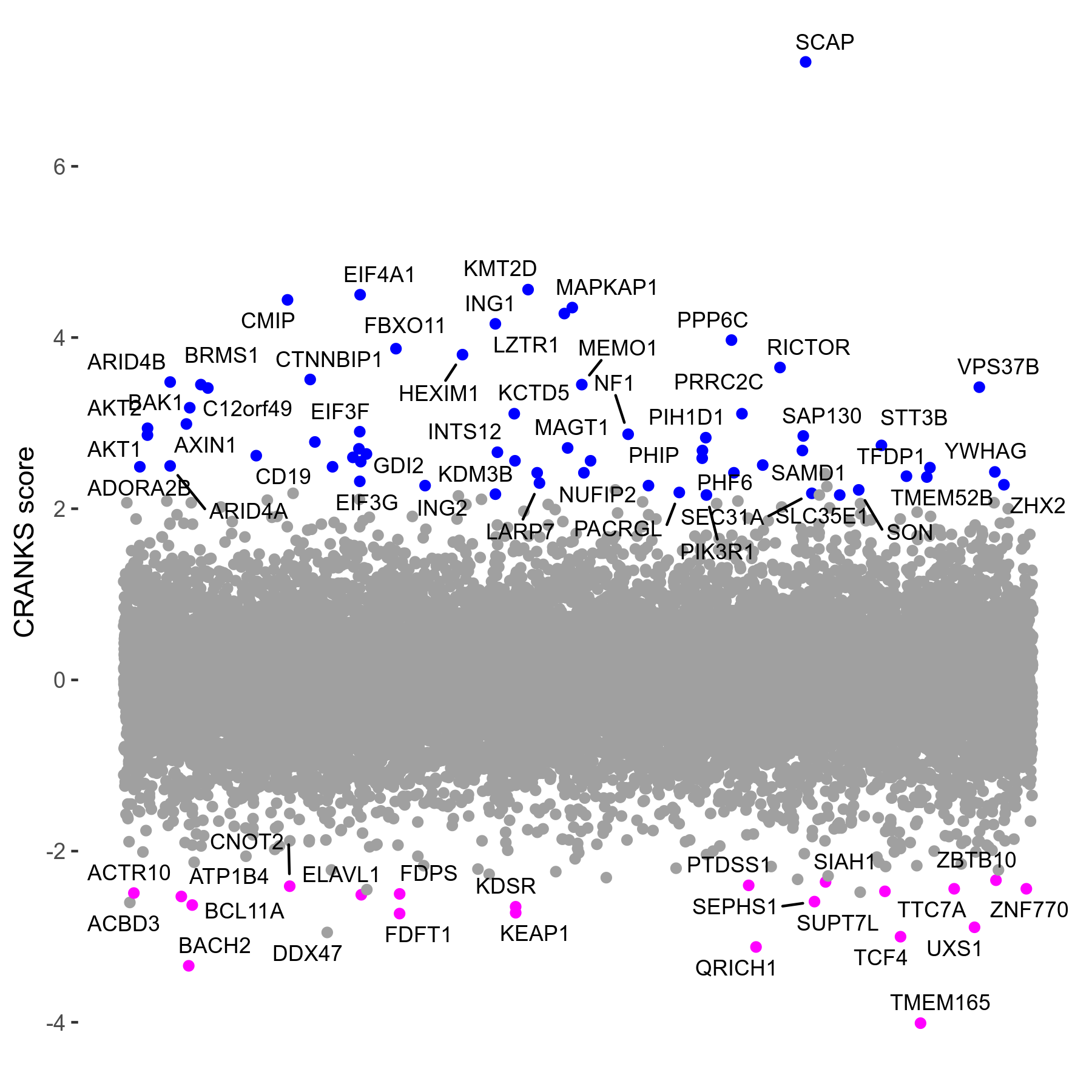
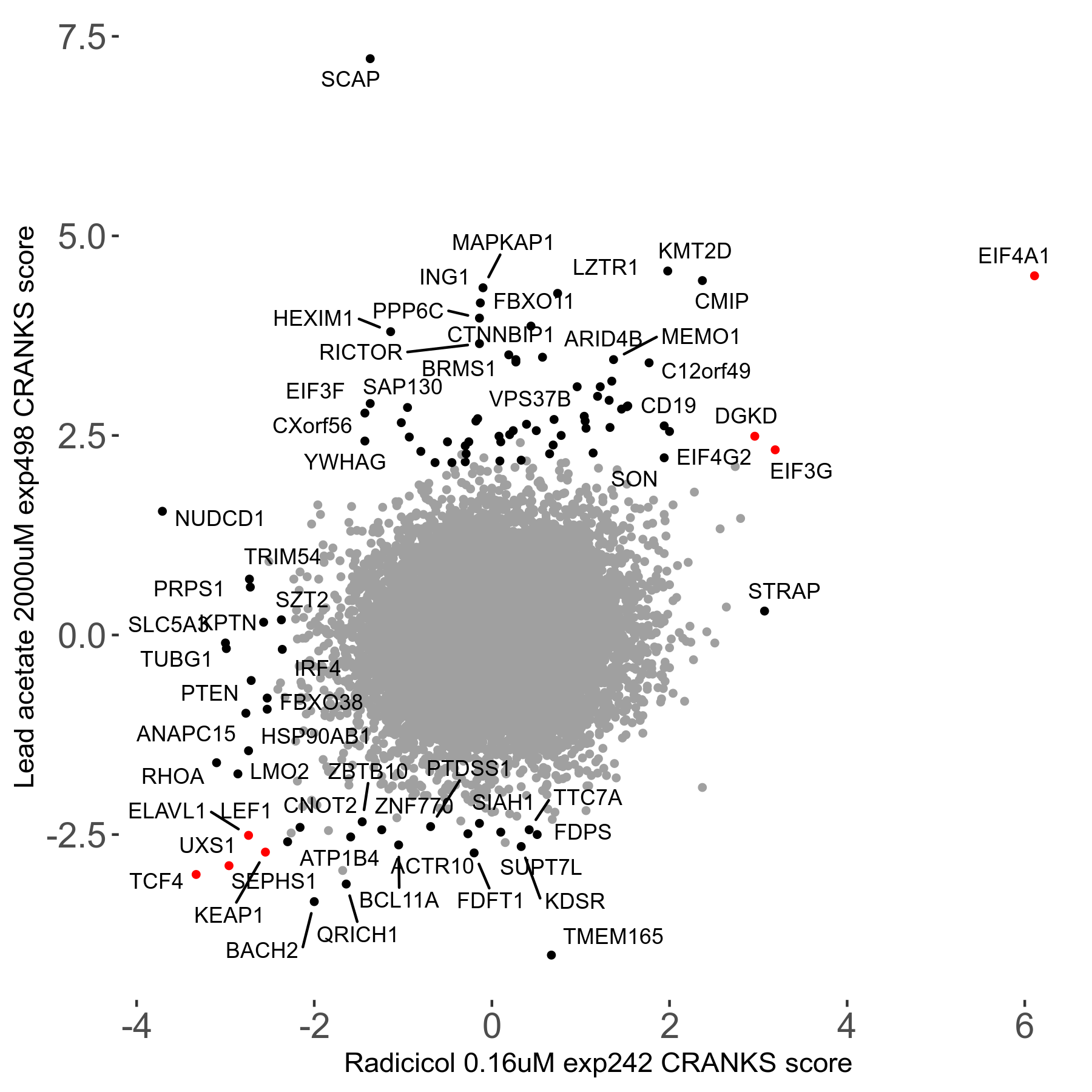
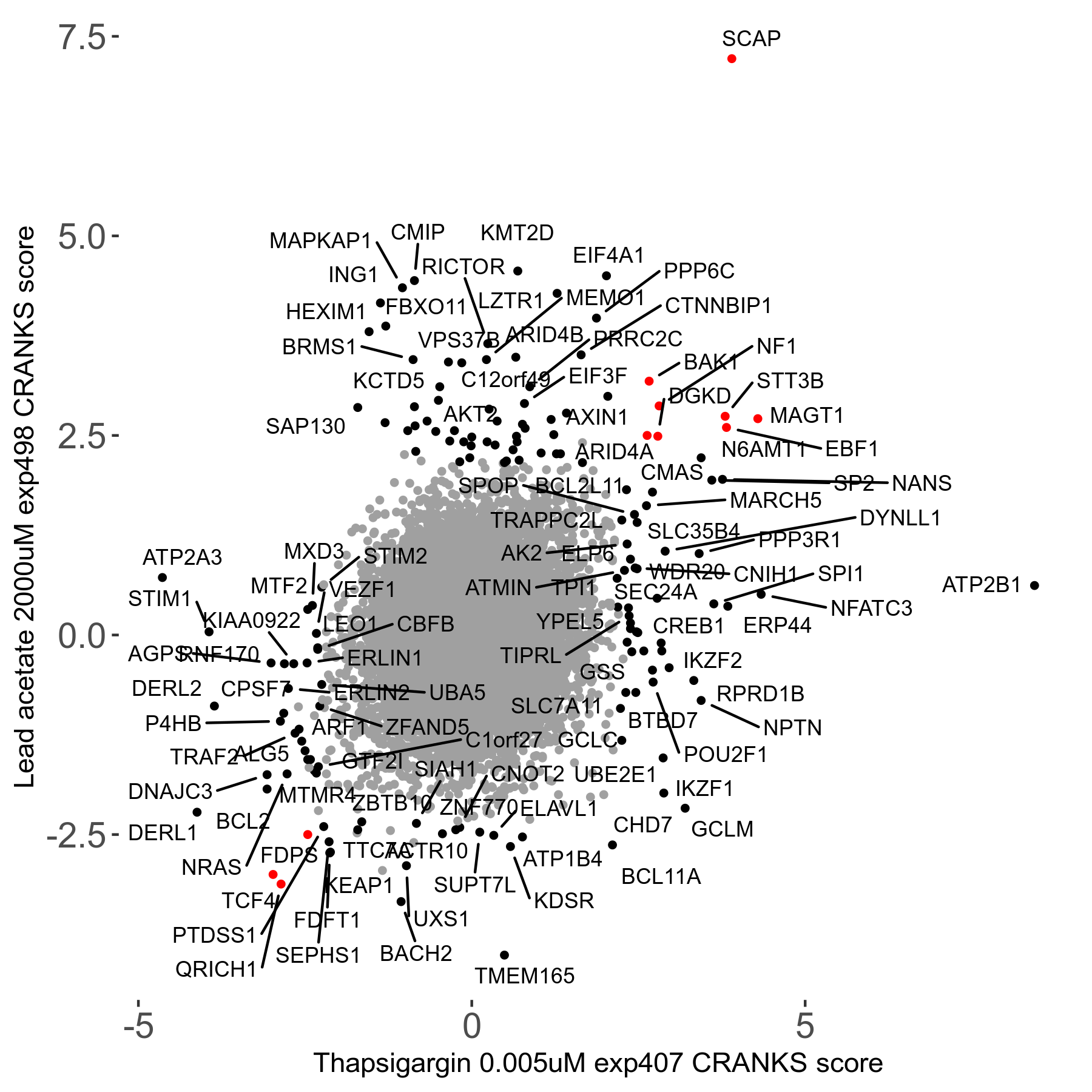
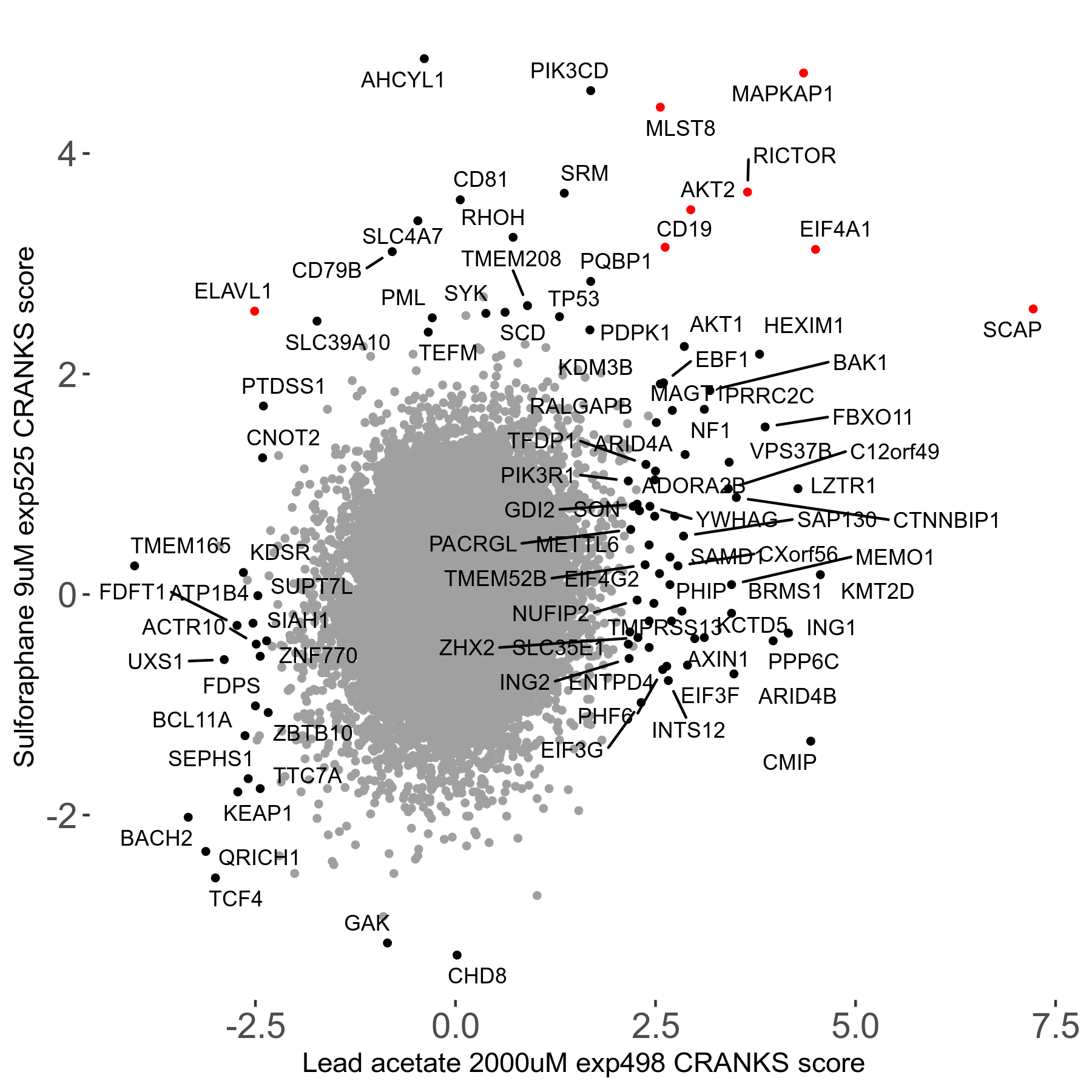
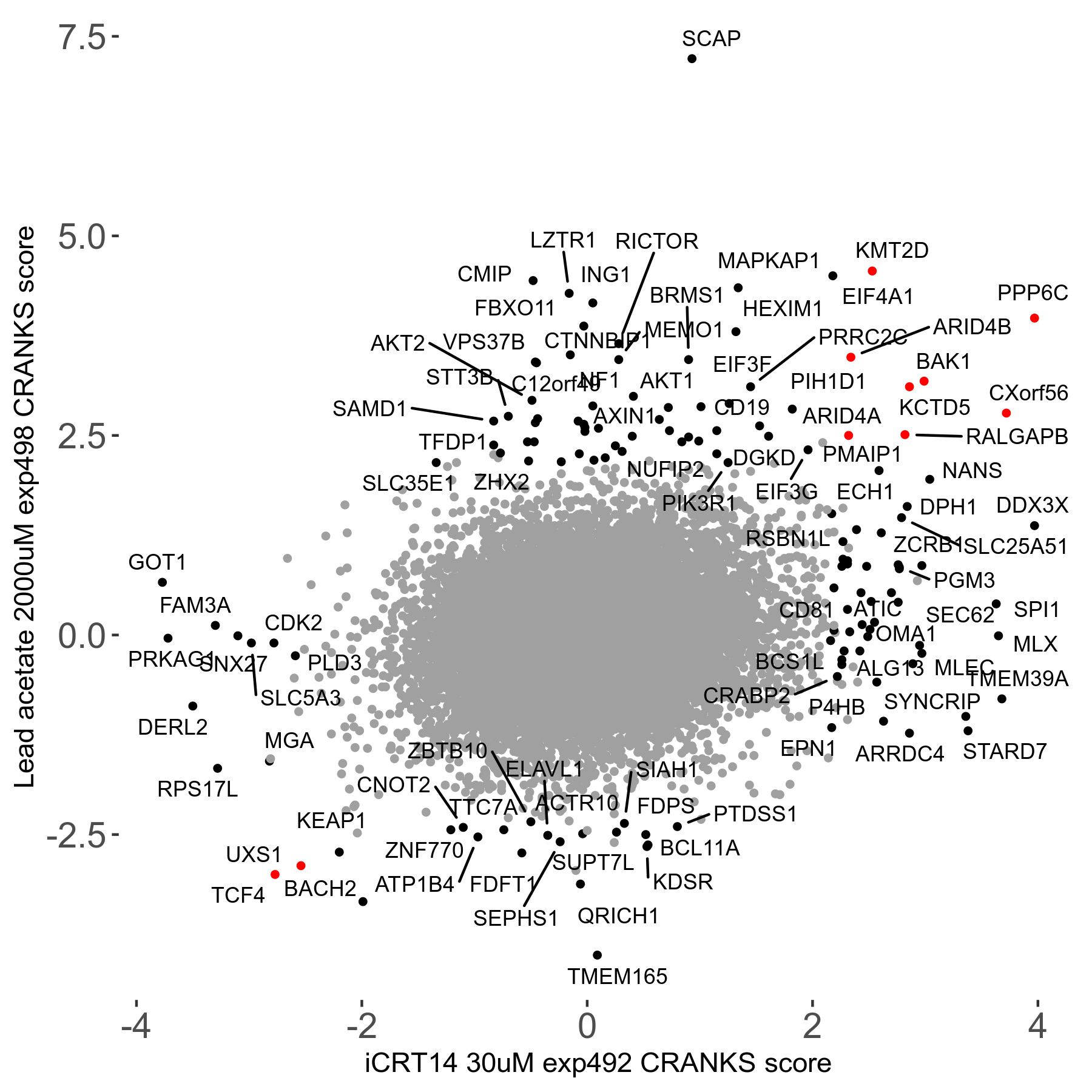
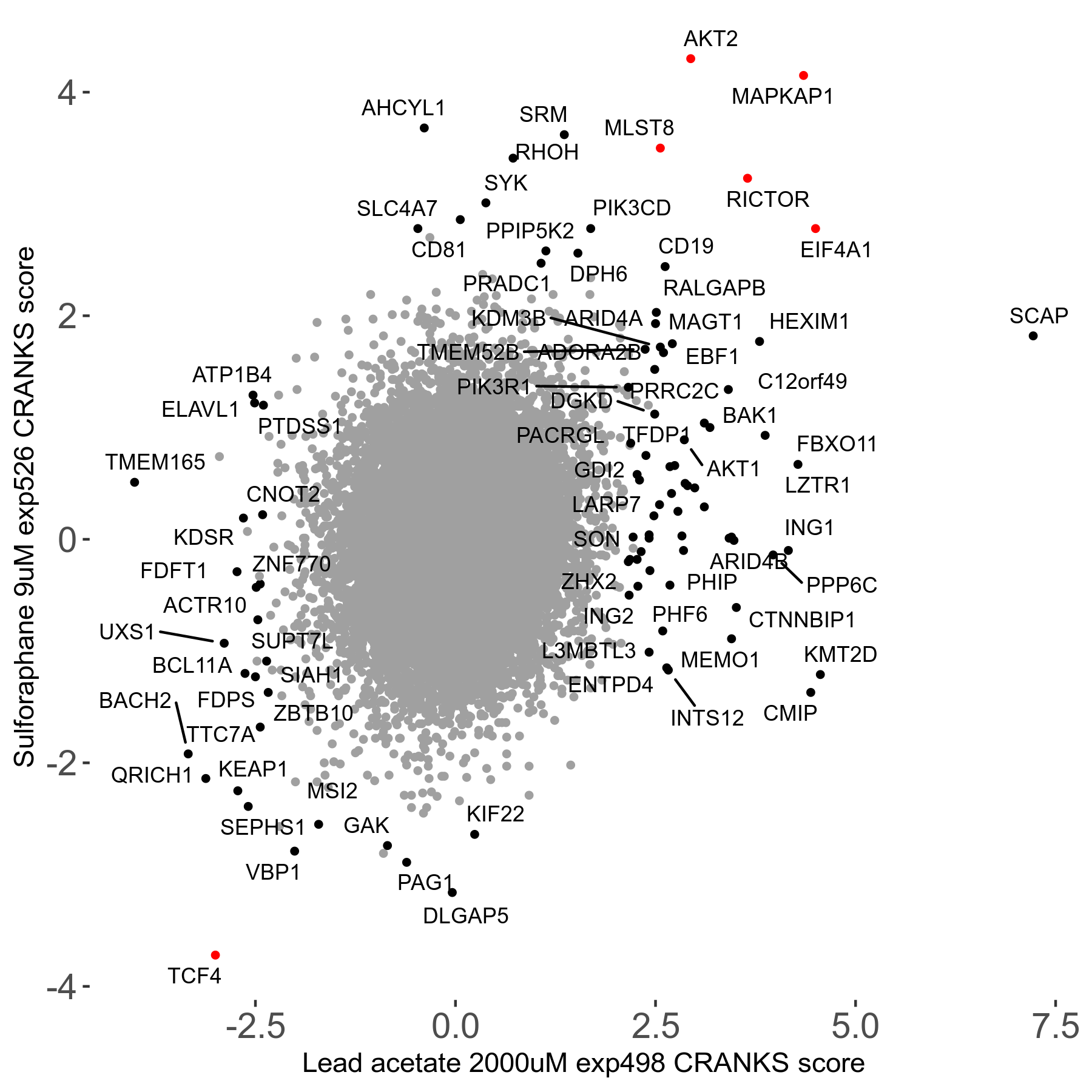
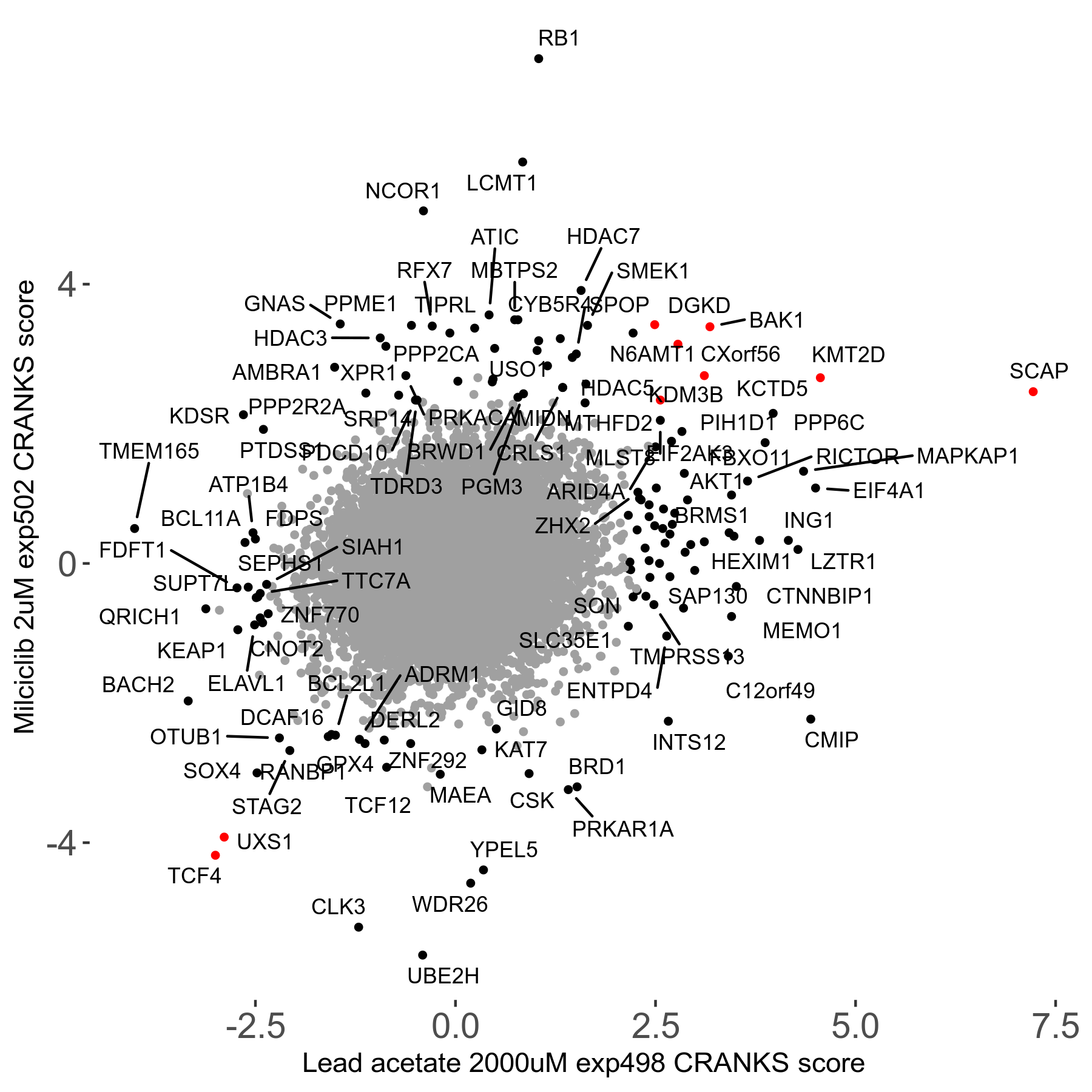
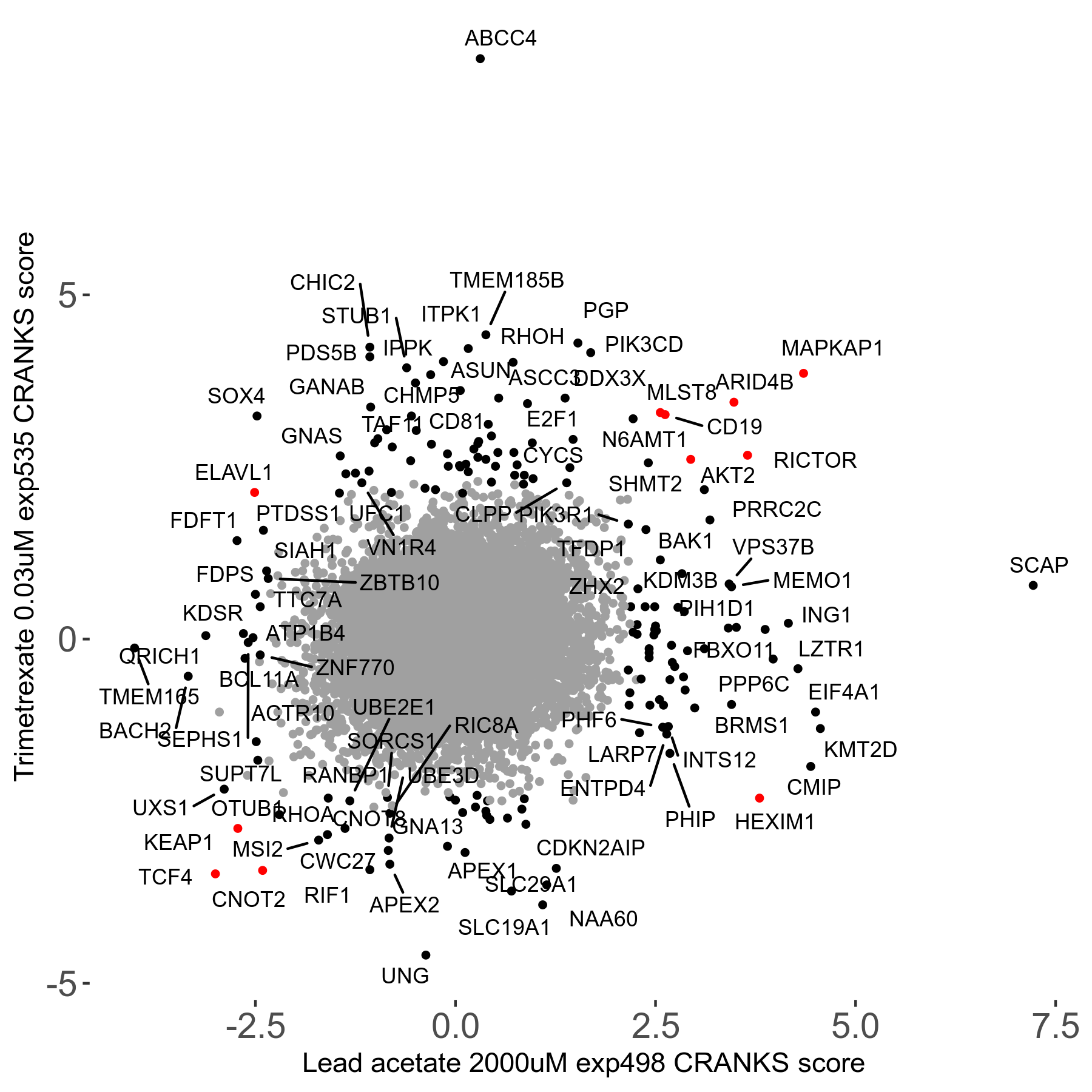
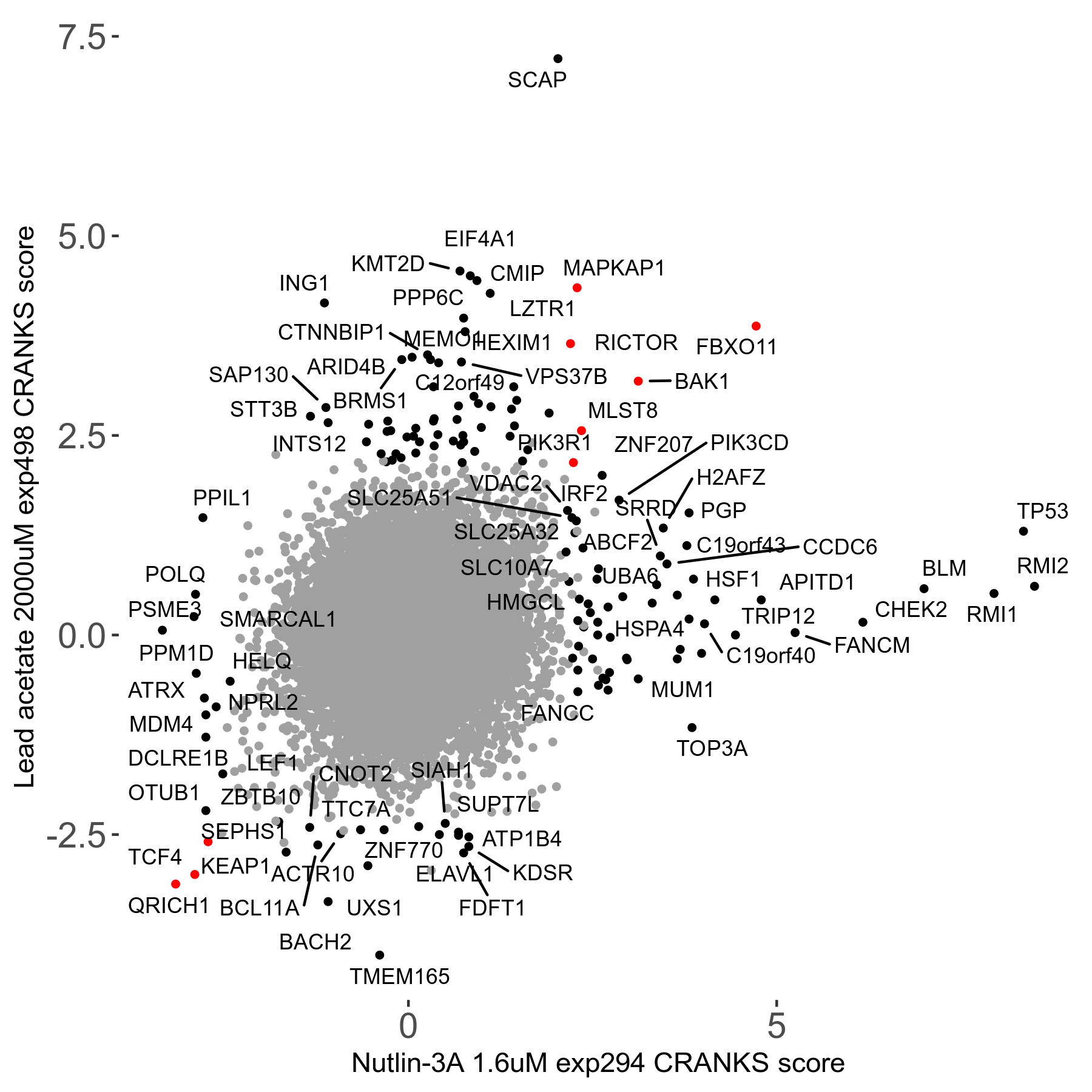
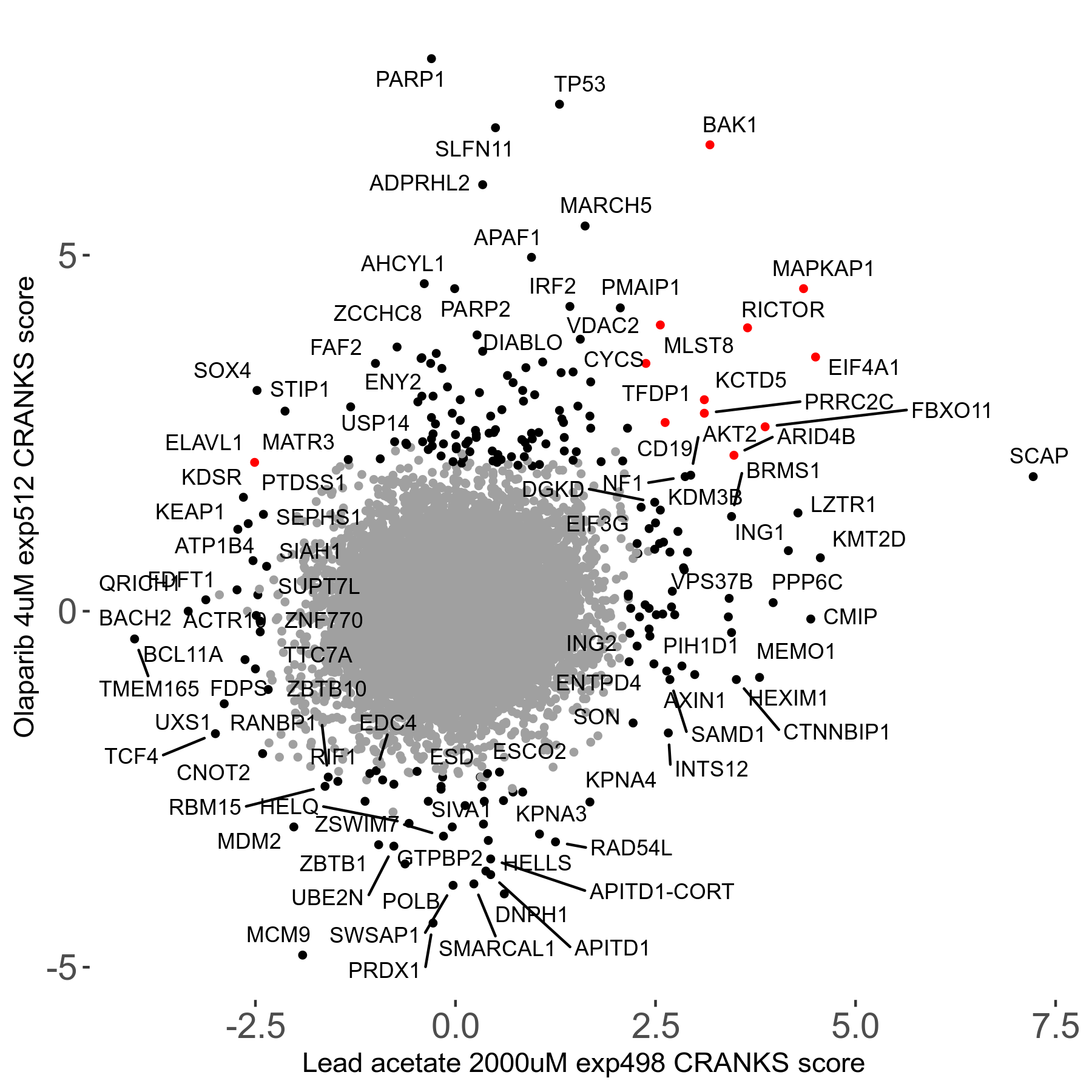
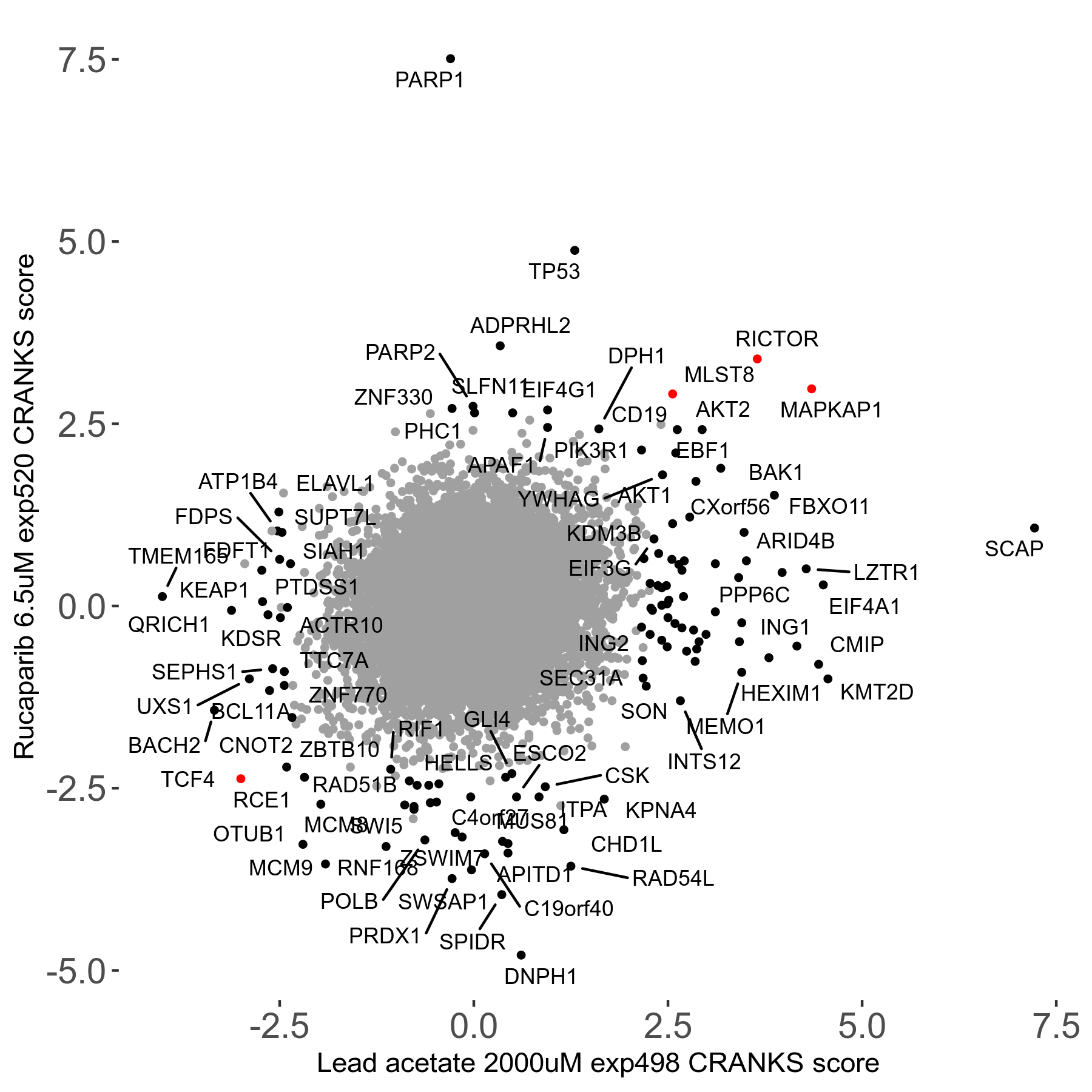
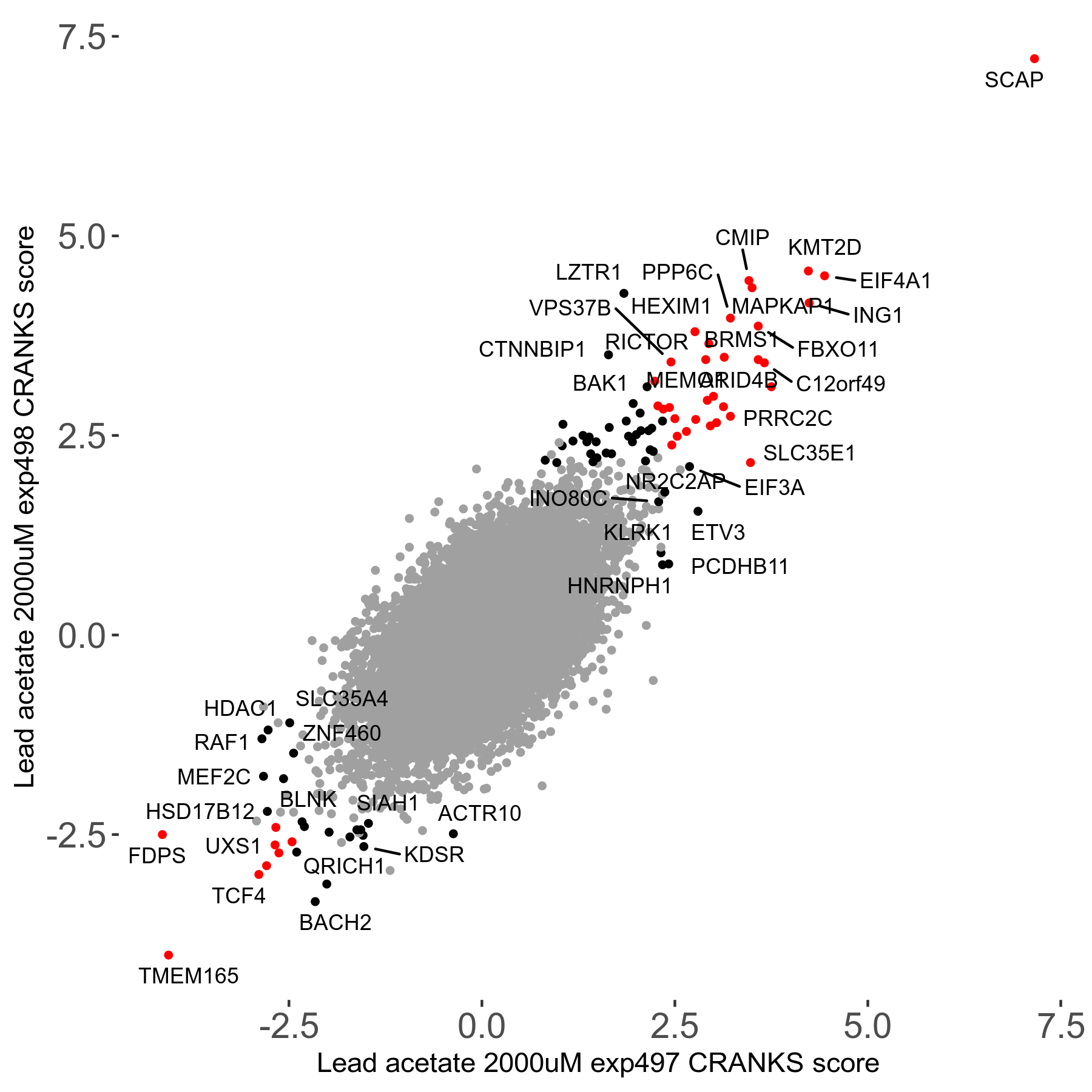
Screen Results
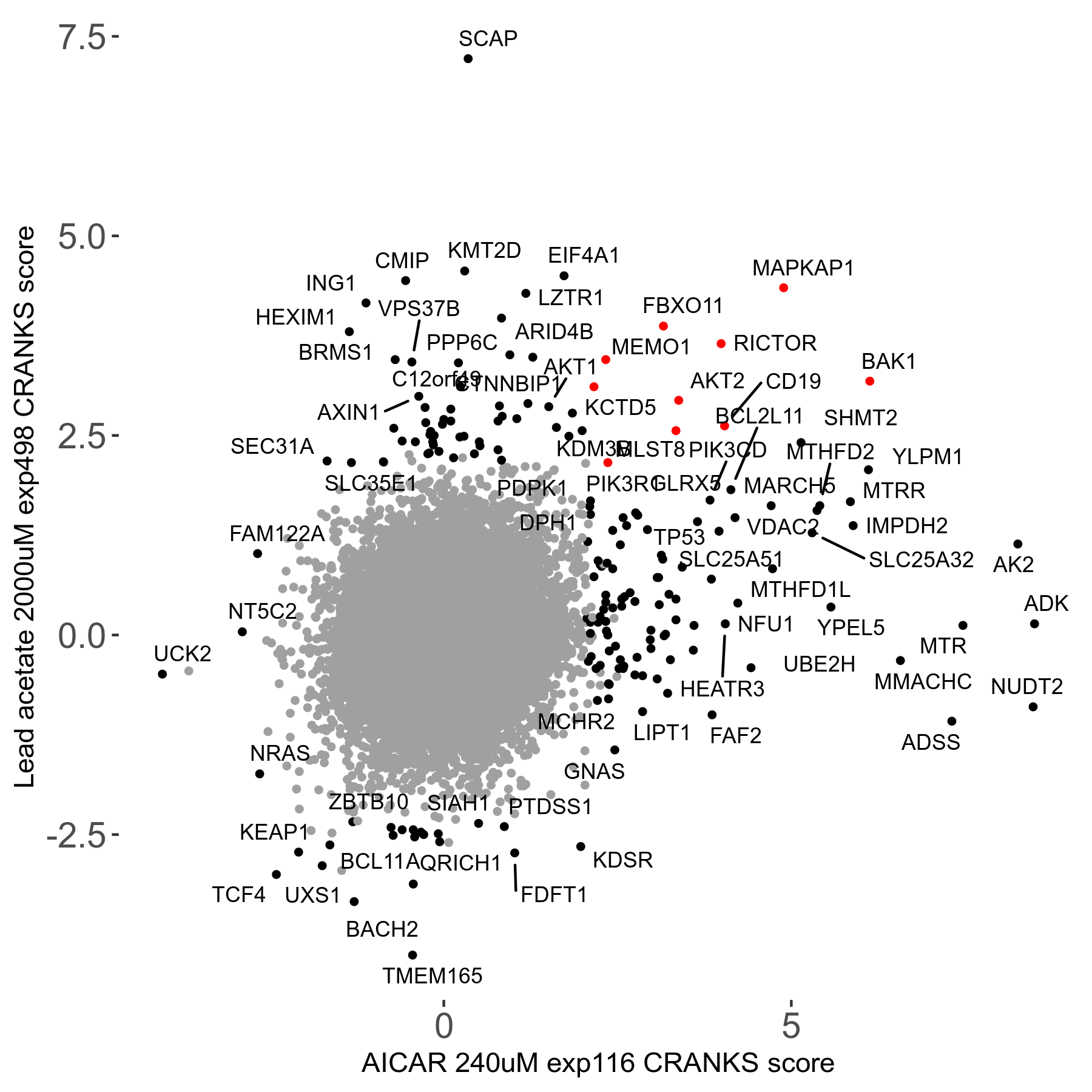
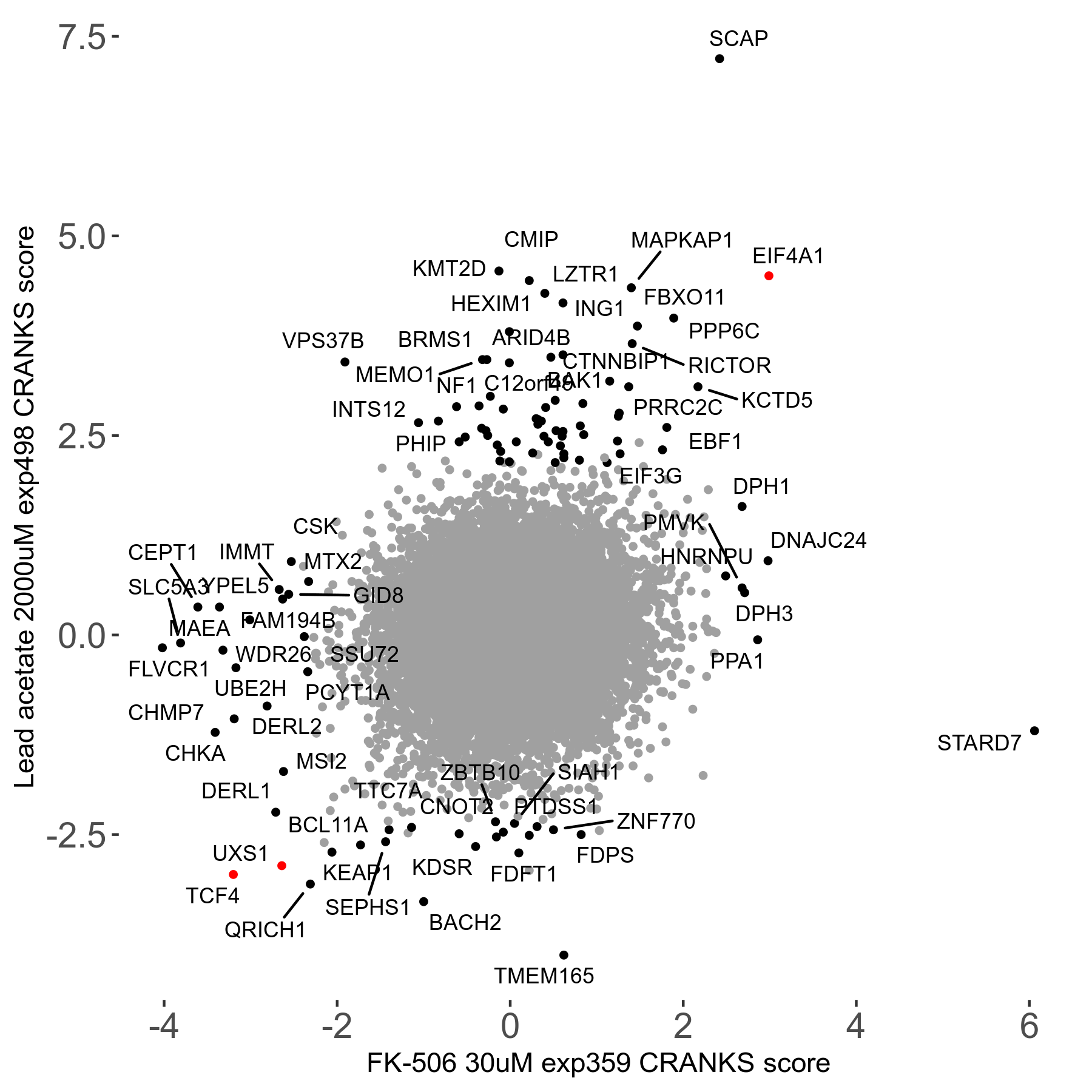
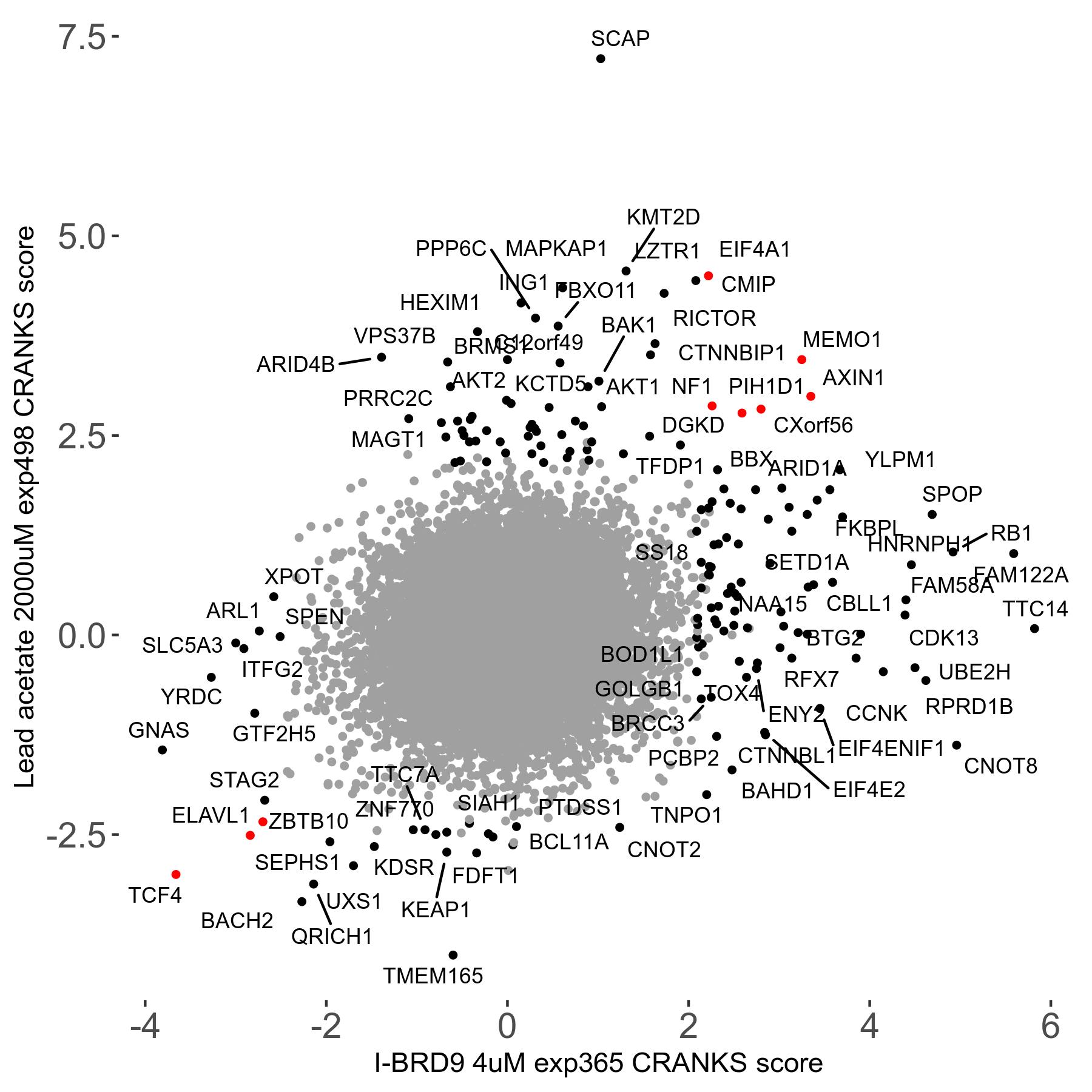
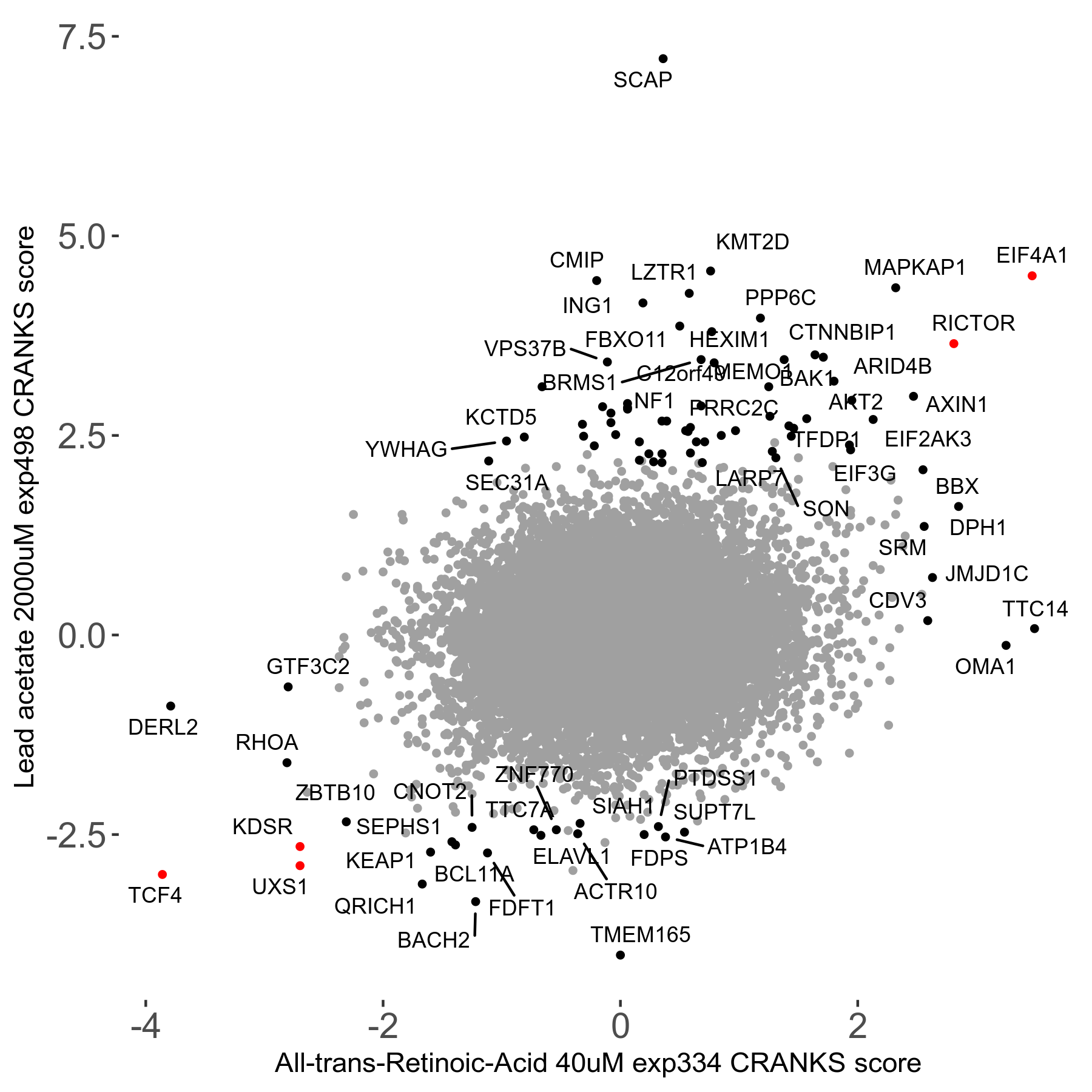
| Sensitive/Resistant hits (FDR<0.05) | CRANKS | Score Plot | Top 30 Genes | Screen Similarity | Top 30 Sensitive GO terms | Top 30 Resistant GO terms |
|---|---|---|---|---|---|---|
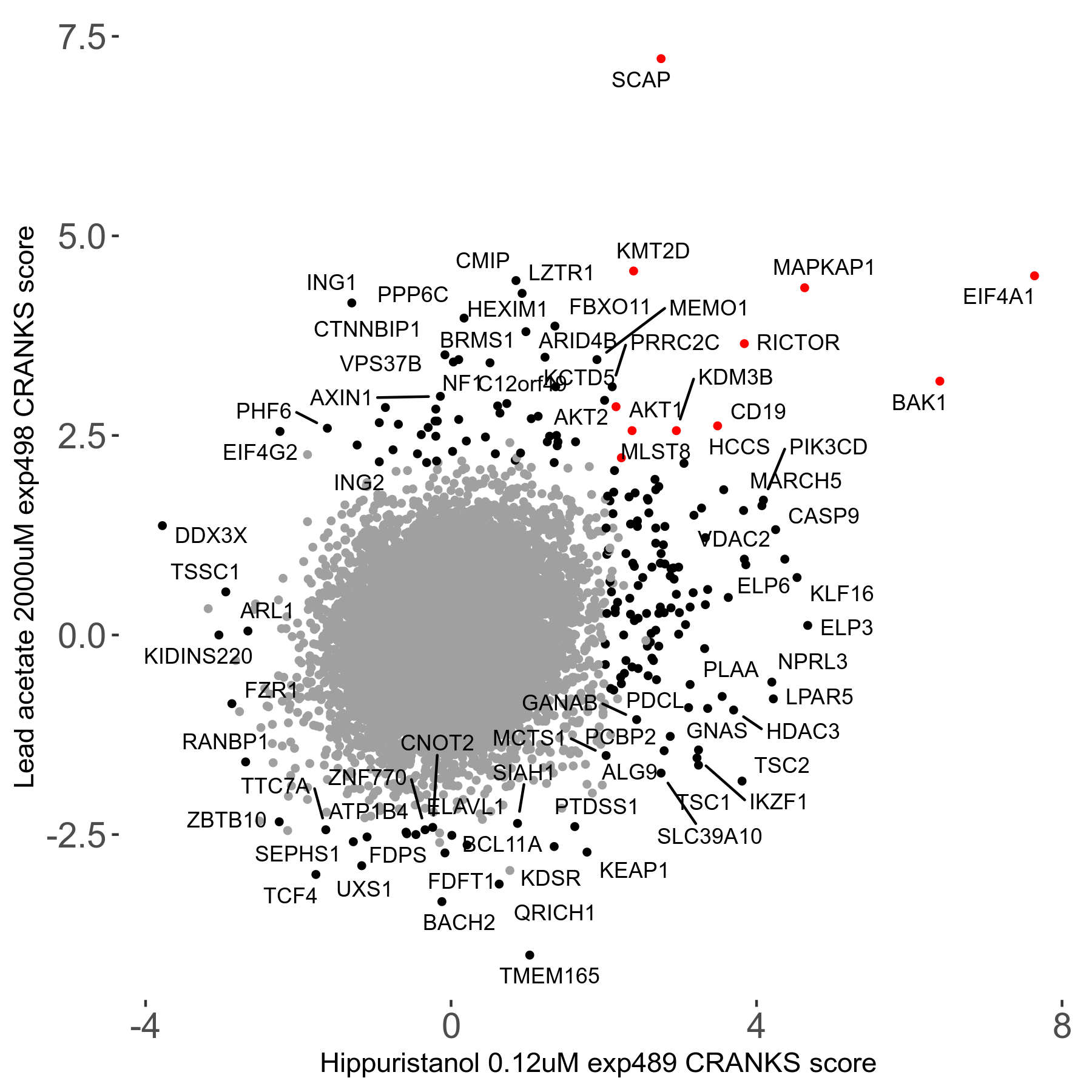
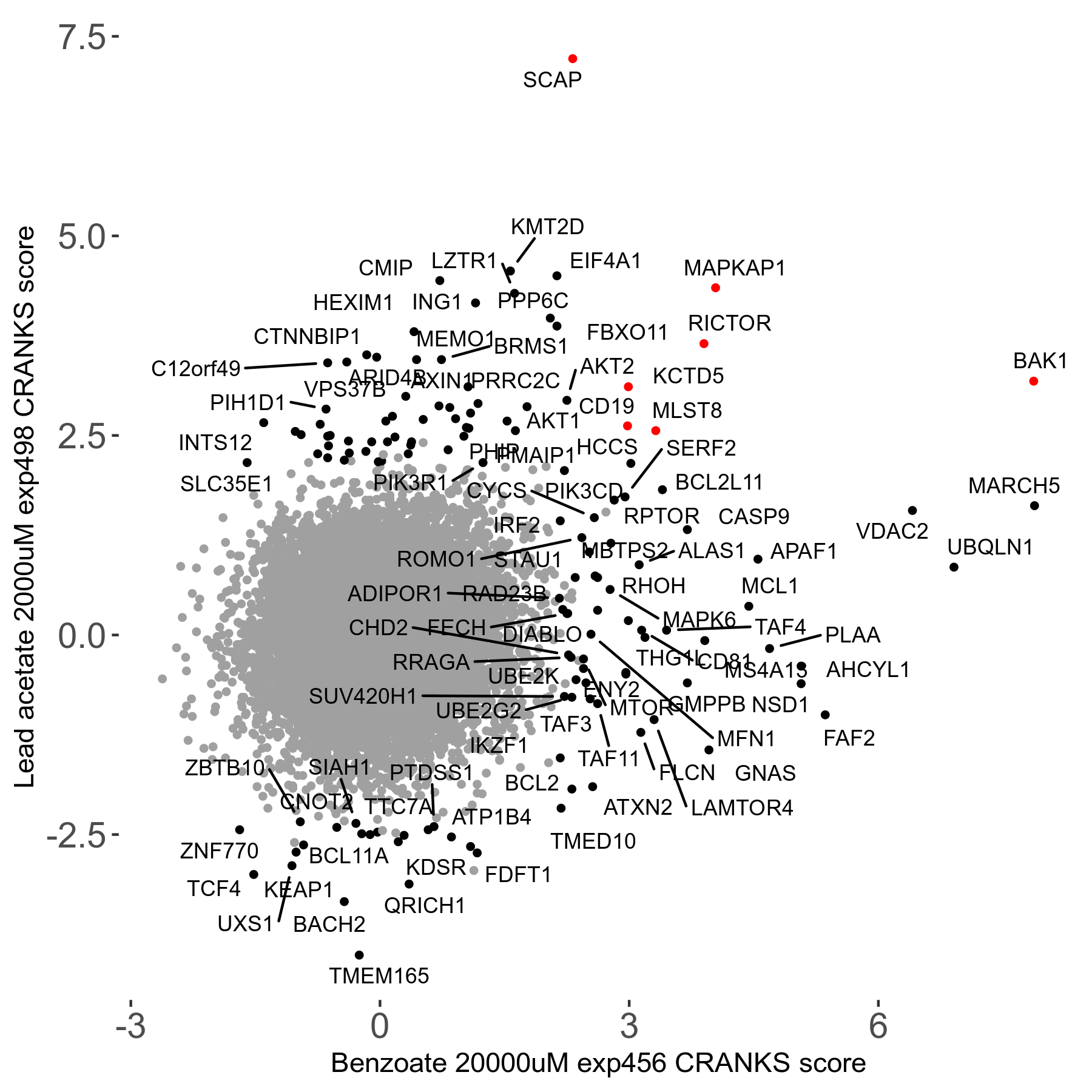
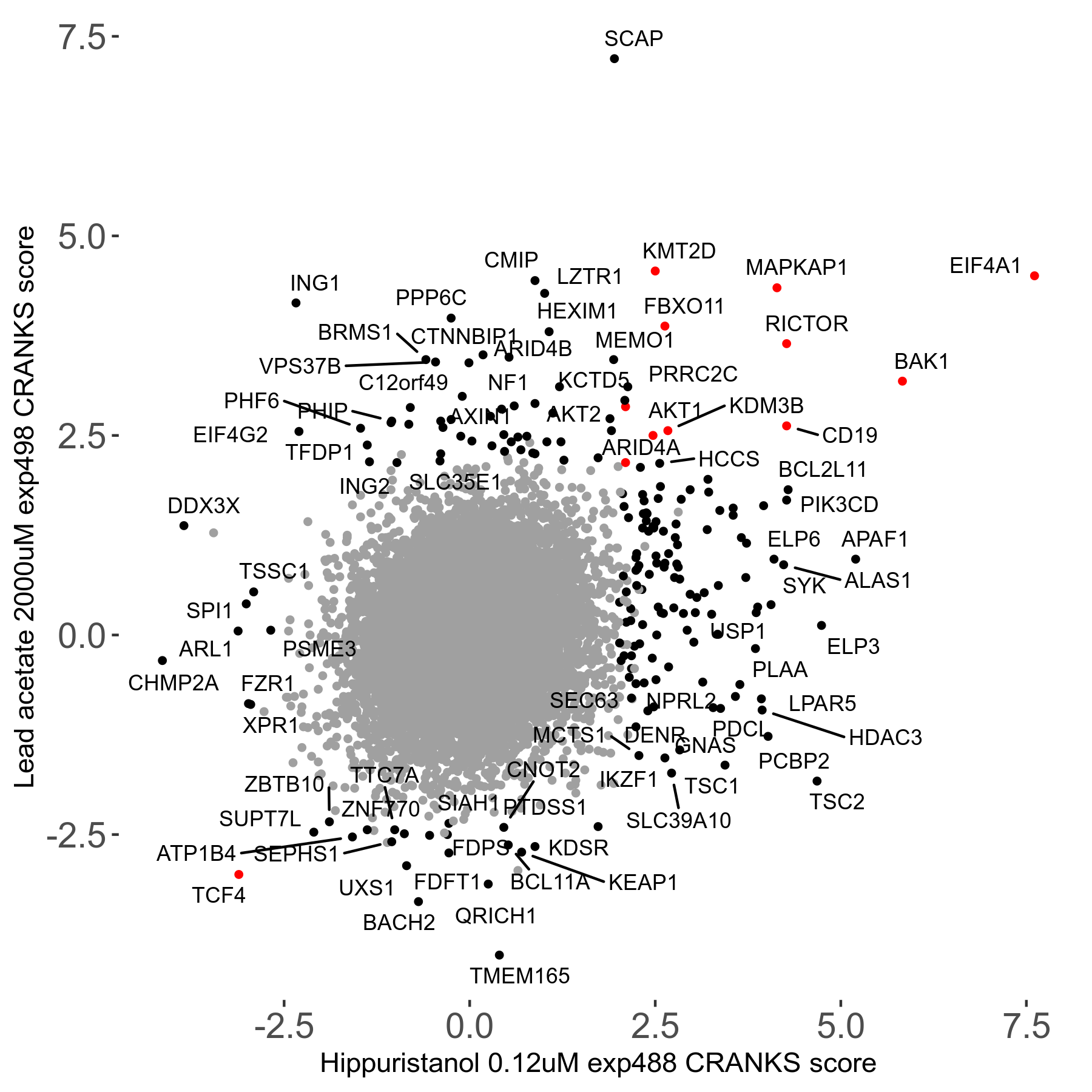
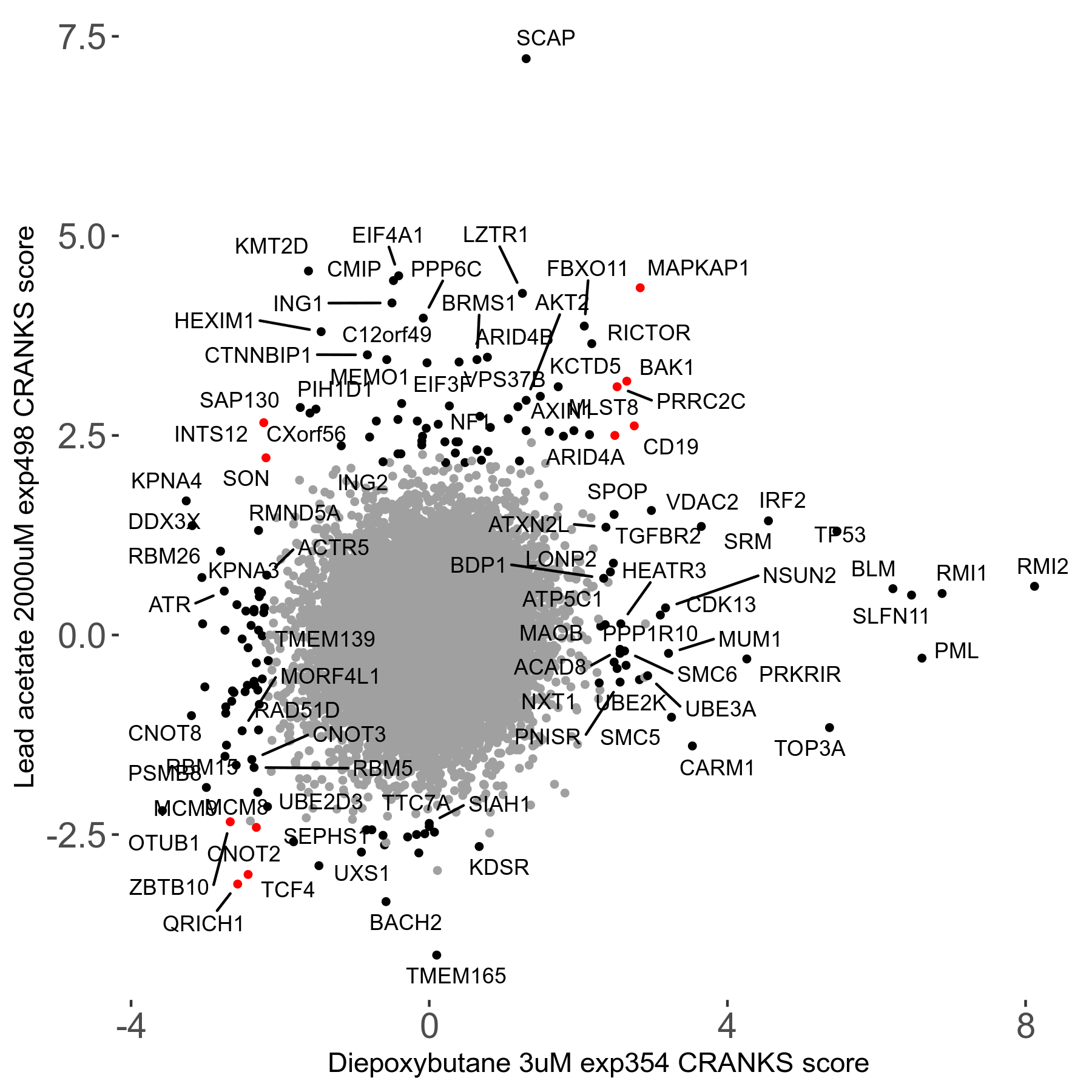
| 21/63 | Scores |