DERL2
Gene Information
- Official Symbol: DERL2
- Official Name: derlin 2
- Aliases and Previous Symbols: N/A
- Entrez ID: 51009
- UniProt: Q9GZP9
- Interactions: BioGRID
- PubMed articles: Open PubMed
- OMIM: Open OMIM
Function Summary
- Entrez Summary: N/A
- UniProt Summary: Functional component of endoplasmic reticulum-associated degradation (ERAD) for misfolded lumenal glycoproteins, but not that of misfolded nonglycoproteins. May act by forming a channel that allows the retrotranslocation of misfolded glycoproteins into the cytosol where they are ubiquitinated and degraded by the proteasome. May mediate the interaction between VCP and misfolded glycoproteins (PubMed:16186509, PubMed:16449189). May also be involved in endoplasmic reticulum stress-induced pre-emptive quality control, a mechanism that selectively attenuates the translocation of newly synthesized proteins into the endoplasmic reticulum and reroutes them to the cytosol for proteasomal degradation (PubMed:26565908). {ECO:0000269|PubMed:16186509, ECO:0000269|PubMed:16449189, ECO:0000269|PubMed:26565908}.
Pfam Domains GO Terms
Pfam Domains
| DER1 |
GO Terms
| signal recognition particle receptor complex |
| signal recognition particle |
| Hrd1p ubiquitin ligase ERAD-L complex |
| negative regulation of retrograde protein transport, ER to cytosol |
| negative regulation of protein exit from endoplasmic reticulum |
| negative regulation of ERAD pathway |
| regulation of retrograde protein transport, ER to cytosol |
| suckling behavior |
| endoplasmic reticulum mannose trimming |
| ubiquitin-specific protease binding |
| endoplasmic reticulum to cytosol transport |
| retrograde protein transport, ER to cytosol |
| protein demannosylation |
| protein alpha-1,2-demannosylation |
| endoplasmic reticulum quality control compartment |
| protein exit from endoplasmic reticulum |
| misfolded protein binding |
| regulation of protein exit from endoplasmic reticulum |
| regulation of ERAD pathway |
| protein deglycosylation |
| negative regulation of intracellular protein transport |
| negative regulation of response to endoplasmic reticulum stress |
| negative regulation of proteasomal protein catabolic process |
| negative regulation of intracellular transport |
| multi-organism behavior |
| ubiquitin-dependent ERAD pathway |
| negative regulation of proteolysis involved in cellular protein catabolic process |
| regulation of response to endoplasmic reticulum stress |
| feeding behavior |
| negative regulation of cellular protein catabolic process |
| ERAD pathway |
| endoplasmic reticulum unfolded protein response |
| integral component of endoplasmic reticulum membrane |
| negative regulation of cellular protein localization |
| late endosome |
| cellular response to unfolded protein |
| negative regulation of protein catabolic process |
| cellular response to topologically incorrect protein |
| response to unfolded protein |
| positive regulation of cell growth |
| negative regulation of protein transport |
| regulation of proteasomal protein catabolic process |
| negative regulation of establishment of protein localization |
| response to topologically incorrect protein |
| regulation of proteolysis involved in cellular protein catabolic process |
| regulation of intracellular protein transport |
| regulation of cellular protein catabolic process |
| negative regulation of cellular catabolic process |
| early endosome |
| response to endoplasmic reticulum stress |
| positive regulation of growth |
| negative regulation of catabolic process |
| proteasome-mediated ubiquitin-dependent protein catabolic process |
| proteasomal protein catabolic process |
| regulation of intracellular transport |
| negative regulation of proteolysis |
| regulation of protein catabolic process |
| glycoprotein metabolic process |
| regulation of cell growth |
| negative regulation of transport |
| ubiquitin-dependent protein catabolic process |
| modification-dependent protein catabolic process |
| regulation of cellular protein localization |
| modification-dependent macromolecule catabolic process |
| behavior |
| proteolysis involved in cellular protein catabolic process |
| cellular protein catabolic process |
| regulation of growth |
| protein catabolic process |
| regulation of protein transport |
| regulation of proteolysis |
| regulation of cellular response to stress |
| regulation of peptide transport |
| regulation of establishment of protein localization |
| regulation of cellular catabolic process |
| cellular macromolecule catabolic process |
| regulation of cellular localization |
| positive regulation of cell population proliferation |
| endoplasmic reticulum membrane |
| intracellular protein transport |
| regulation of catabolic process |
| response to organonitrogen compound |
| endoplasmic reticulum |
| regulation of protein localization |
| carbohydrate derivative metabolic process |
| negative regulation of cellular protein metabolic process |
| macromolecule catabolic process |
| organonitrogen compound catabolic process |
| response to nitrogen compound |
| negative regulation of protein metabolic process |
| proteolysis |
| regulation of response to stress |
| protein transport |
| intracellular transport |
| peptide transport |
| amide transport |
| cellular protein localization |
| cellular macromolecule localization |
| establishment of protein localization |
| regulation of cell population proliferation |
| negative regulation of response to stimulus |
| cellular response to stress |
| organic substance catabolic process |
| cellular catabolic process |
| establishment of localization in cell |
| nitrogen compound transport |
| regulation of transport |
| membrane |
CRISPR Data
Compound Hit Most Correlated Genes in Chemogenomics Tissues where Essential in the Avana Dataset (DepMap 20Q1)
Compound Hit
Most Correlated Genes in Chemogenomics
| Gene | Correlation |
|---|---|
| UBE2J1 | 0.602 |
| MANF | 0.602 |
| DNAJC3 | 0.563 |
| UBE2G2 | 0.504 |
| ALG5 | 0.499 |
| CCDC101 | 0.496 |
| DERL1 | 0.492 |
| ENY2 | 0.468 |
| GMPPB | 0.468 |
| XPO6 | 0.464 |
| GTF3C2 | 0.453 |
| CBFB | 0.436 |
| SLC35B1 | 0.435 |
| DNAJB11 | 0.43 |
| GTF2I | 0.426 |
| MED16 | 0.421 |
| DPAGT1 | 0.421 |
| EIF4E | 0.418 |
| NXT1 | 0.418 |
| MAPK14 | 0.417 |
| RNMT | 0.414 |
| TMED10 | 0.411 |
| RPTOR | 0.411 |
| C1orf27 | 0.41 |
| VMP1 | 0.409 |
| DAD1 | 0.403 |
| MAX | 0.402 |
| TMEM41B | 0.401 |
Tissues where Essential in the Avana Dataset (DepMap 20Q1)
Global Fraction of Cell Lines Where Essential: 55/739
| Tissue | Fraction Of Cell Lines Where Essential |
|---|---|
| 1290807.0 | 1/1 |
| 909776.0 | 0/1 |
| bile duct | 1/28 |
| blood | 5/28 |
| bone | 2/26 |
| breast | 2/33 |
| central nervous system | 3/56 |
| cervix | 0/4 |
| colorectal | 0/17 |
| esophagus | 0/13 |
| fibroblast | 0/1 |
| gastric | 3/16 |
| kidney | 3/21 |
| liver | 4/20 |
| lung | 3/75 |
| lymphocyte | 3/16 |
| ovary | 1/26 |
| pancreas | 1/24 |
| peripheral nervous system | 1/16 |
| plasma cell | 2/15 |
| prostate | 0/1 |
| skin | 2/24 |
| soft tissue | 2/9 |
| thyroid | 0/2 |
| upper aerodigestive | 3/22 |
| urinary tract | 0/29 |
| uterus | 0/5 |
Essentiality in NALM6
- Essentiality Rank: 1850
- Expression level (log2 read counts): 5.74
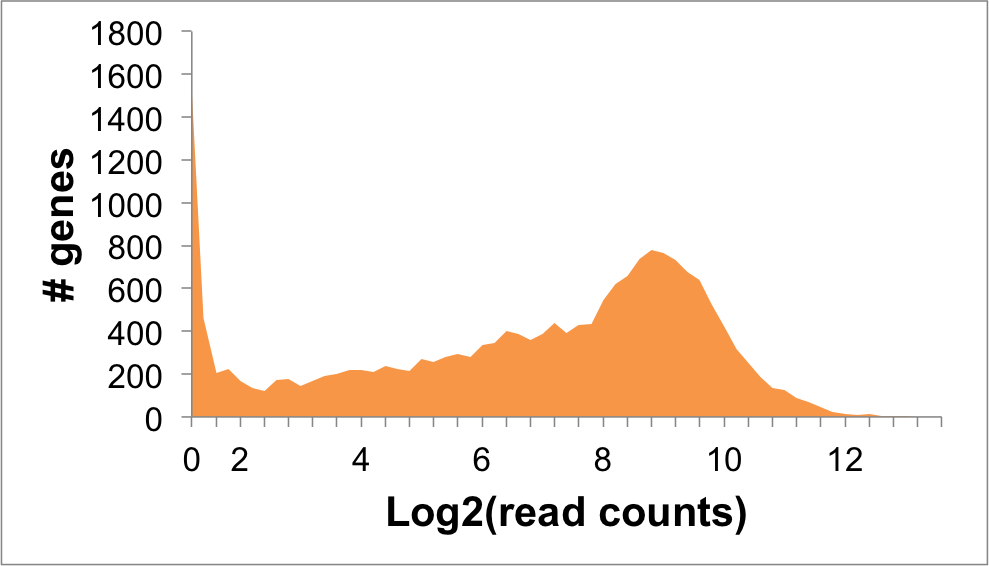
Expression Distribution
DERL2 Expression in NALM6 Cells: 5.74