RTCB
Gene Information
- Official Symbol: RTCB
- Official Name: RNA 2',3'-cyclic phosphate and 5'-OH ligase
- Aliases and Previous Symbols: N/A
- Entrez ID: 51493
- UniProt: Q9Y3I0
- Interactions: BioGRID
- PubMed articles: Open PubMed
- OMIM: Open OMIM
Function Summary
- Entrez Summary: N/A
- UniProt Summary: Catalytic subunit of the tRNA-splicing ligase complex that acts by directly joining spliced tRNA halves to mature-sized tRNAs by incorporating the precursor-derived splice junction phosphate into the mature tRNA as a canonical 3',5'- phosphodiester. May act as an RNA ligase with broad substrate specificity, and may function toward other RNAs. {ECO:0000269|PubMed:21311021, ECO:0000269|PubMed:24870230}.
Pfam Domains GO Terms
Pfam Domains
| UPF0027 |
GO Terms
| RNA ligase (ATP) activity |
| tRNA exon ligation |
| tRNA exon ligation utilizing 2,3 cyclic phosphate of 5-exon as source of linkage phosphate |
| RNA exon ligation |
| tRNA-splicing ligase complex |
| vinculin binding |
| tRNA splicing, via endonucleolytic cleavage and ligation |
| RNA splicing, via endonucleolytic cleavage and ligation |
| tRNA processing |
| placenta development |
| nuclear envelope |
| tRNA metabolic process |
| in utero embryonic development |
| ncRNA processing |
| RNA splicing |
| reproductive structure development |
| reproductive system development |
| ncRNA metabolic process |
| chordate embryonic development |
| embryo development ending in birth or egg hatching |
| developmental process involved in reproduction |
| intracellular membrane-bounded organelle |
| RNA processing |
| endoplasmic reticulum membrane |
| embryo development |
| RNA binding |
| reproductive process |
| reproduction |
| ATP binding |
| RNA metabolic process |
| gene expression |
CRISPR Data
Compound Hit Most Correlated Genes in Chemogenomics Tissues where Essential in the Avana Dataset (DepMap 20Q1)
Compound Hit
Most Correlated Genes in Chemogenomics
| Gene | Correlation |
|---|---|
| THG1L | 0.718 |
| DPAGT1 | 0.704 |
| GFPT1 | 0.69 |
| UAP1 | 0.658 |
| RPTOR | 0.648 |
| GMPPB | 0.645 |
| DOHH | 0.636 |
| ALG1 | 0.632 |
| GTF3C3 | 0.63 |
| RNMT | 0.628 |
| TRIAP1 | 0.626 |
| GNB1L | 0.618 |
| TRMT5 | 0.615 |
| PMM2 | 0.614 |
| RPP21 | 0.613 |
| OSGEP | 0.613 |
| ATXN10 | 0.611 |
| ADSL | 0.609 |
| NAE1 | 0.608 |
| TSEN34 | 0.608 |
| TTI2 | 0.608 |
| DHPS | 0.604 |
| SRP14 | 0.602 |
| EIF4E | 0.601 |
| POLR3H | 0.593 |
| YARS | 0.592 |
| EXOC4 | 0.589 |
| CCDC101 | 0.589 |
| MED16 | 0.588 |
| GTF3C2 | 0.585 |
| UHRF1 | 0.581 |
| CCT7 | 0.572 |
| HSPD1 | 0.569 |
| GTF3C4 | 0.567 |
| ALG14 | 0.566 |
| AARS | 0.566 |
| ALG2 | 0.564 |
| ALG13 | 0.564 |
| PELO | 0.561 |
| GTF3C6 | 0.558 |
| PPP4C | 0.558 |
| DDX1 | 0.552 |
| VRK1 | 0.551 |
| DDX20 | 0.551 |
| TPRKB | 0.545 |
| PPCS | 0.544 |
| TELO2 | 0.54 |
| CSNK2B | 0.538 |
| RPE | 0.536 |
| ENY2 | 0.534 |
| NARS | 0.532 |
| TBCA | 0.532 |
| POLR3B | 0.529 |
| MEAF6 | 0.529 |
| TTI1 | 0.529 |
| HARS | 0.527 |
| MED4 | 0.526 |
| CRCP | 0.524 |
| TSEN54 | 0.523 |
| EPRS | 0.52 |
| RABGGTB | 0.519 |
| GEMIN8 | 0.519 |
| VEZT | 0.519 |
| TGS1 | 0.518 |
| TRNT1 | 0.517 |
| DOLK | 0.517 |
| PLAA | 0.516 |
| CTC1 | 0.515 |
| MLLT1 | 0.513 |
| ZRSR2 | 0.513 |
| ELP3 | 0.512 |
| TEN1 | 0.512 |
| ATP5J | 0.512 |
| DDX55 | 0.511 |
| GEMIN5 | 0.51 |
| ALG5 | 0.509 |
| DPF2 | 0.508 |
| RFT1 | 0.507 |
| CMTR1 | 0.507 |
| ORAOV1 | 0.506 |
| TAZ | 0.505 |
| GEMIN6 | 0.503 |
| UXT | 0.502 |
| UTP23 | 0.502 |
| DR1 | 0.5 |
| CEP57 | 0.5 |
| LSM11 | 0.498 |
| RPN1 | 0.496 |
| RBM14 | 0.494 |
| HAUS8 | 0.493 |
| HCCS | 0.491 |
| FNTA | 0.491 |
| PAM16 | 0.491 |
| RAD50 | 0.49 |
| C15orf41 | 0.489 |
| DDOST | 0.488 |
| DHX29 | 0.486 |
| CBFB | 0.485 |
| TAMM41 | 0.485 |
| WDR59 | 0.485 |
| C14orf166 | 0.484 |
| TMEM41B | 0.484 |
| UBE2G2 | 0.482 |
| SSB | 0.481 |
| TBCB | 0.48 |
| SRP68 | 0.48 |
| BAP1 | 0.479 |
| SEH1L | 0.479 |
| NSMCE1 | 0.478 |
| PPA2 | 0.478 |
| CASP9 | 0.478 |
| MARS | 0.477 |
| MTOR | 0.476 |
| UBA3 | 0.473 |
| APAF1 | 0.473 |
| WDR24 | 0.473 |
| PYROXD1 | 0.471 |
| DICER1 | 0.47 |
| ATP5B | 0.47 |
| ENO1 | 0.47 |
| LAMTOR5 | 0.47 |
| VMP1 | 0.469 |
| URM1 | 0.469 |
| DHDDS | 0.467 |
| CTDNEP1 | 0.467 |
| MAPK14 | 0.467 |
| VHL | 0.466 |
| RAD1 | 0.466 |
| DAD1 | 0.465 |
| ELP2 | 0.465 |
| BUB3 | 0.464 |
| TIMM13 | 0.463 |
| PAX5 | 0.463 |
| SNAPC5 | 0.462 |
| PPP6C | 0.462 |
| EXOC3 | 0.461 |
| OTUD5 | 0.458 |
| ZNF608 | 0.457 |
| SLC25A26 | 0.457 |
| FH | 0.456 |
| EEF1E1 | 0.455 |
| UBE2K | 0.455 |
| PIK3R4 | 0.454 |
| ELAC2 | 0.452 |
| SRP72 | 0.451 |
| SPCS2 | 0.448 |
| TTC4 | 0.448 |
| UBA5 | 0.446 |
| DCTN6 | 0.445 |
| OIP5 | 0.444 |
| PGGT1B | 0.444 |
| PPP2R3C | 0.444 |
| GTF3C5 | 0.443 |
| TSEN2 | 0.443 |
| CARS | 0.44 |
| ATP5O | 0.439 |
| TRMT10A | 0.439 |
| TIMM22 | 0.439 |
| SLC35B1 | 0.438 |
| SUPV3L1 | 0.437 |
| TXNL4B | 0.436 |
| TBP | 0.436 |
| EIF1AX | 0.436 |
| FAF2 | 0.434 |
| WDR82 | 0.434 |
| DCTN5 | 0.433 |
| ALG6 | 0.433 |
| NBN | 0.432 |
| THOC6 | 0.432 |
| SEC62 | 0.431 |
| DIABLO | 0.43 |
| ATIC | 0.43 |
| XRCC3 | 0.429 |
| TP53RK | 0.426 |
| PDAP1 | 0.425 |
| BAK1 | 0.424 |
| RPIA | 0.424 |
| SNF8 | 0.423 |
| ROMO1 | 0.423 |
| NSD1 | 0.423 |
| ZNF259 | 0.422 |
| MAD2L1 | 0.422 |
| EIF3F | 0.421 |
| NUP50 | 0.421 |
| WDR92 | 0.42 |
| TKT | 0.419 |
| NAA20 | 0.418 |
| SRPRB | 0.417 |
| MED23 | 0.416 |
| THOC3 | 0.416 |
| LSMD1 | 0.415 |
| SEC61G | 0.415 |
| NHLRC2 | 0.415 |
| ACTR10 | 0.414 |
| METAP2 | 0.414 |
| TAF1A | 0.413 |
| SLC25A19 | 0.412 |
| RFK | 0.412 |
| AASDHPPT | 0.411 |
| CHCHD4 | 0.411 |
| TMED10 | 0.41 |
| LAMTOR4 | 0.41 |
| LSM10 | 0.41 |
| GTF2A2 | 0.408 |
| FBXW7 | 0.408 |
| TCF4 | 0.408 |
| ATP2A2 | 0.407 |
| SSBP2 | 0.407 |
| UBE2L3 | 0.407 |
| TPK1 | 0.404 |
| CINP | 0.404 |
| CRLS1 | 0.404 |
| WDR1 | 0.403 |
| ATM | 0.403 |
| SOD1 | 0.403 |
| WARS | 0.403 |
| CIAO1 | 0.403 |
| SRP19 | 0.402 |
| SDHA | 0.402 |
| ATP5A1 | 0.402 |
| CCDC115 | 0.402 |
Tissues where Essential in the Avana Dataset (DepMap 20Q1)
Global Fraction of Cell Lines Where Essential: 422/739
| Tissue | Fraction Of Cell Lines Where Essential |
|---|---|
| 1290807.0 | 0/1 |
| 909776.0 | 0/1 |
| bile duct | 14/28 |
| blood | 12/28 |
| bone | 13/26 |
| breast | 13/33 |
| central nervous system | 39/56 |
| cervix | 1/4 |
| colorectal | 9/17 |
| esophagus | 9/13 |
| fibroblast | 0/1 |
| gastric | 8/16 |
| kidney | 12/21 |
| liver | 12/20 |
| lung | 47/75 |
| lymphocyte | 6/16 |
| ovary | 15/26 |
| pancreas | 15/24 |
| peripheral nervous system | 8/16 |
| plasma cell | 5/15 |
| prostate | 0/1 |
| skin | 11/24 |
| soft tissue | 6/9 |
| thyroid | 2/2 |
| upper aerodigestive | 15/22 |
| urinary tract | 21/29 |
| uterus | 3/5 |
Essentiality in NALM6
- Essentiality Rank: 925
- Expression level (log2 read counts): 6.76
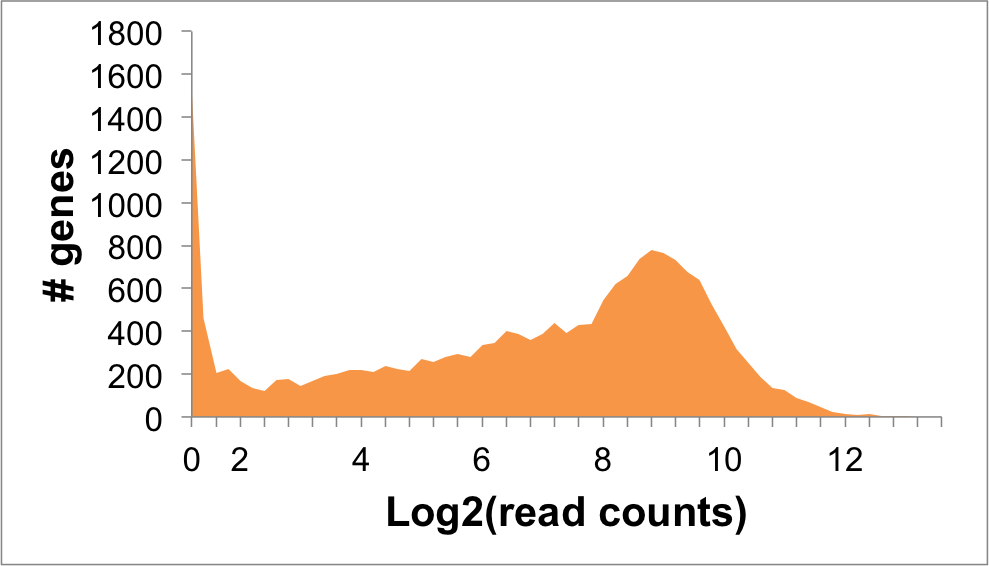
Expression Distribution
RTCB Expression in NALM6 Cells: 6.76